Les données obtenues via le GBIF
| Dans ce module, vous découvrirez les données primaires sur la biodiversité et les principes que le GBIF suit en matière de données. Vous apprendrez comment le GBIF rend les données primaires sur la biodiversité accessibles, quels sont les types de données acceptés et comment le GBIF utilise l’épine dorsale taxonomique pour fournir des informations taxonomiques. Vous aurez également l’occasion de passer en revue les différentes métriques disponibles pour les données du portail. |
Données primaires sur la biodiversité
|
Dans cette vidéo (12:03), Cecilie Svenningsen, chef de produit des données du GBIF, explique comment les données sont partagées avec le GBIF et comment elles peuvent être utilisées, tout en décrivant les normes de biodiversité et les types d’ensembles de données du GBIF. Si vous ne pouvez pas regarder la vidéo Vimeo intégrée, vous pouvez la télécharger localement (MP4 - 73,1 Mo). |
Transcription de la présentation
Cliquez pour développer

Diapositive 1 - Comment les données sont-elles partagées sur GBIF.org ?
Il convient de garder à l’esprit que les éditeurs de données du GBIF possèdent et conservent leurs données, qu’ils partagent ensuite avec le GBIF.
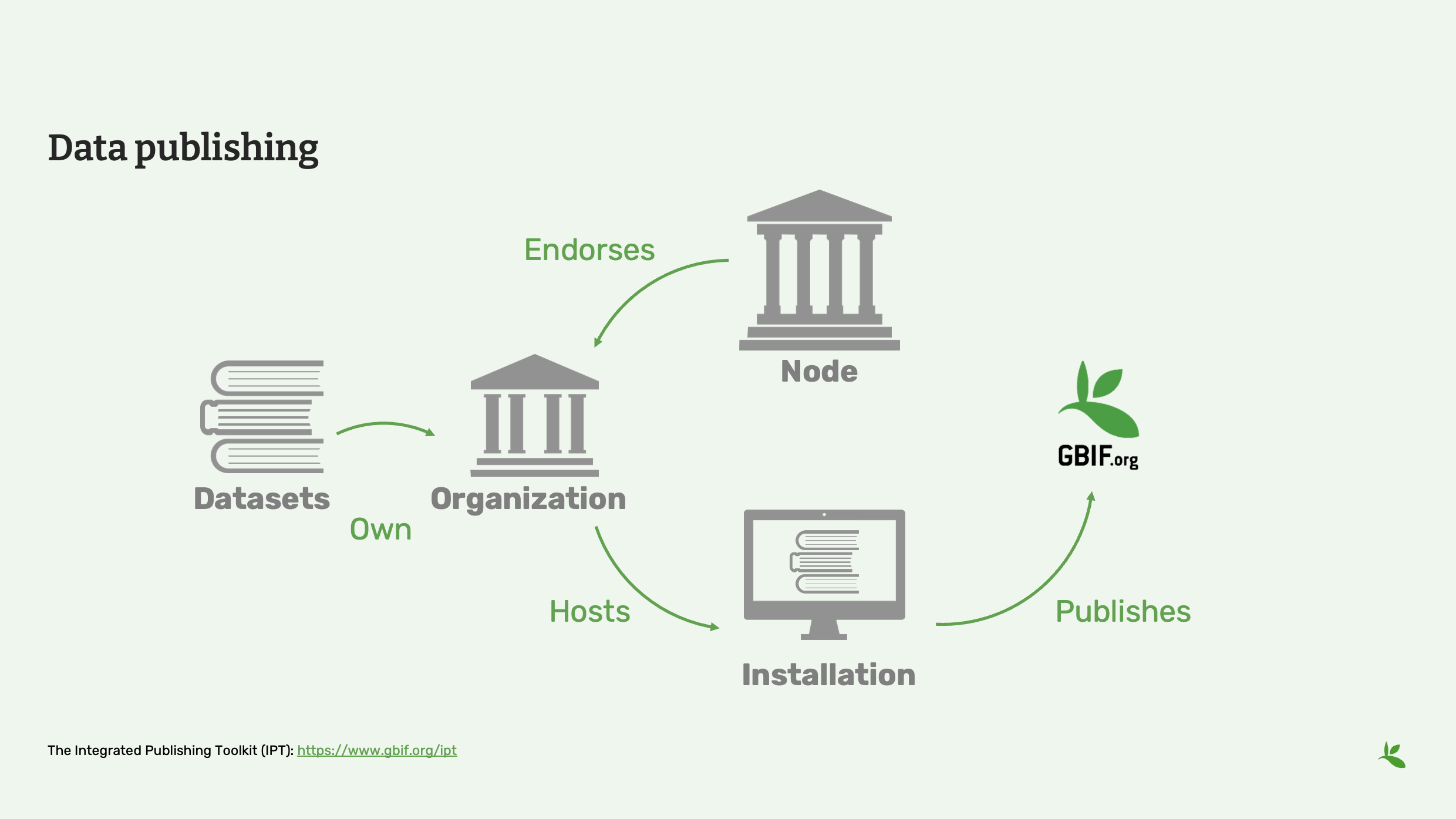
Diapositive 2 - Publication de données
Le fonctionnement est donc le suivant : nous avons un ensemble de données qui appartient à une organisation. Cette organisation doit être approuvée par un nœud du GBIF, qui peut être un nœud national ou tout autre nœud participant, pour pouvoir partager des données avec le GBIF.
Ces données doivent ensuite être hébergées quelque part dans une installation, par exemple la boîte à outils de publication intégrée fournie par le GBIF.
Les ensembles de données y sont donc hébergés, puis ils peuvent être publiés dans le GBIF et partagés avec vous.

Diapositive 3 - Les données partagées sur le GBIF sont hétérogènes
Une autre chose dont il faut tenir compte est que les données du GBIF sont très hétérogènes. Les organisations qui partagent des données avec le GBIF proviennent de sources très diverses, et il s’agit d’un processus approuvé par la communauté, comme je l’ai mentionné, où les nœuds approuvent les organisations de publication, qui peuvent être des universités, des musées, des agences gouvernementales, etc.
Ce qu’ils partagent peut donc provenir de projets de recherche ou d’efforts de suivi, de données issues de la science citoyenne ou de collectes.
Il faut également garder à l’esprit que ces données ne sont pas disponibles en direct sur le GBIF.
Vous pouvez avoir des données qui font partie d’un effort mobilisé d’une collection, d’une collection de musée qui pourrait numériser ses données d’il y a 50 ans, 70 ans, et le jour suivant, elles sont disponibles sur GBIF.
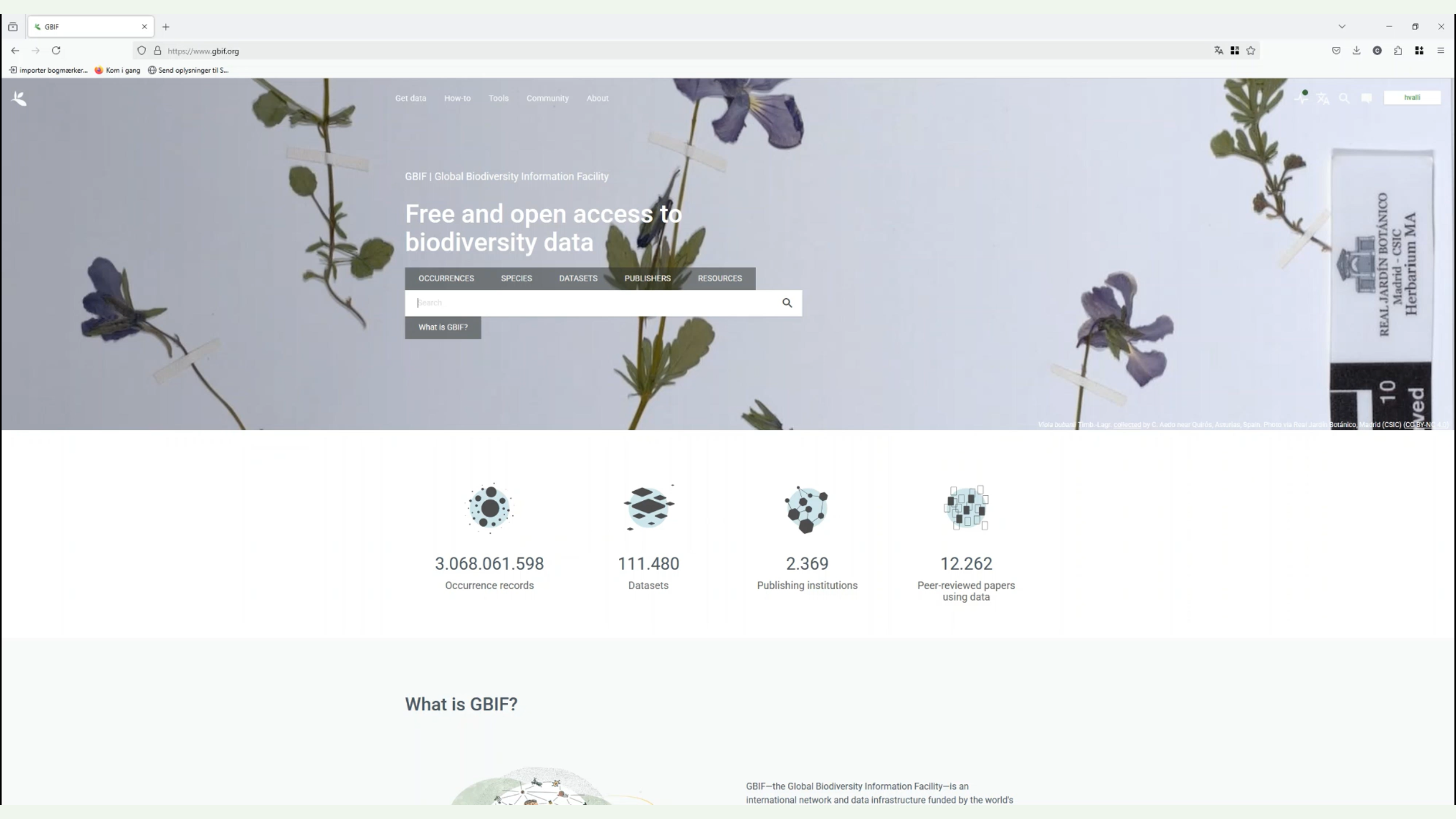
Diapositive 4 - Accéder aux données et les télécharger
Pour vous montrer comment une telle recherche agrégée peut se présenter à vous en tant qu’utilisateur, si vous allez voir une espèce, dans ce cas, je vous montre le papillon à pointe orange. Si, comme la plupart de nos utilisateurs, vous vous rendez sur notre portail et recherchez l’espèce, la première chose que vous verrez sera la page des espèces.
Vous obtiendrez ainsi une description générale de l’espèce et la répartition des occurrences du GBIF. Mais en général, vous devriez aller à la page d’enregistrement des occurrences, où vous pourriez appliquer d’autres filtres.
Dans ce cas, je dis que je ne veux que des entrées datant de 2020 à 2022, je pense. Je précise également qu’ils doivent se situer sur le continent européen.
J’ai donc réduit ma recherche et je clique sur télécharger, où je dois choisir différentes options. Dans ce cas, je choisis l’option simple.
Mais vous pouvez également voir d’autres informations, telles que les problèmes connus pour les données, ce qui pourrait être quelque chose que vous devriez examiner si vous voulez appliquer des filtres supplémentaires pour votre téléchargement, si vous ne voulez pas que ces données soient incluses.
Ainsi, dans votre profil d’utilisateur GBIF, vous pouvez voir tous les téléchargements que vous avez créés.
Dans ce cas, je vous montre celui que je viens de créer, les filtres. Mais vous verrez aussi que 927 ensembles de données ont contribué à cette recherche agrégée.
Les sources de données sont donc très variées. C’est important car cela a un impact sur ce qui sera réellement dans les données que vous téléchargez.
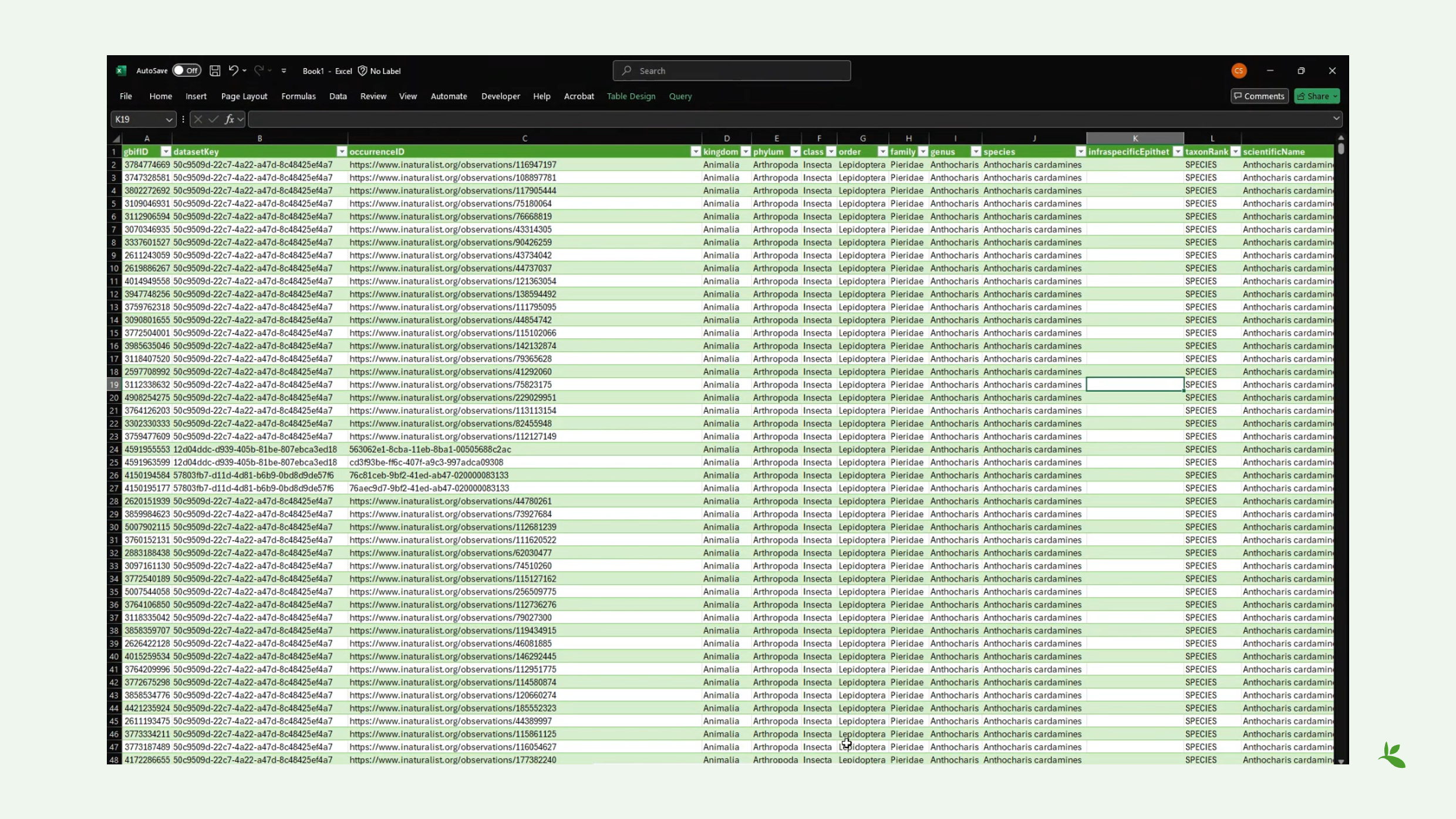
Diapositive 5 - Examen d’un téléchargement
Voici donc un exemple du téléchargement simple que je viens de créer, dans lequel il y a moins de champs disponibles, moins de colonnes. Mais si vous téléchargez l’archive Darwin Core complète, vous disposerez de nombreux champs pour effectuer des recherches. Mais vous remarquerez que certains de ces champs semblent être vides, c’est-à-dire qu’il n’y a pas de valeurs. Mais comme vous pouvez le voir, comme je vous le montre maintenant, vous avez en fait des informations sur la localité partagées par certains éditeurs.
Cela signifie que certains éditeurs peuvent partager leurs données et d’autres non. Et cela peut avoir une incidence sur votre analyse.
Ainsi, par exemple, le compte individuel, qui peut vous donner une idée de l’abondance de l’espèce, n’est pas nécessairement rempli par certains éditeurs, mais l’est par d’autres.
Et comme je l’ai mentionné, si vous téléchargez l’archive Darwin Core complète, où vous avez beaucoup plus de colonnes, vous verrez un ensemble de données beaucoup plus inégal, ce qui a une incidence sur la façon dont vous post-traitez vos données.

Diapositive 6 - Dans quelle mesure les champs que vous téléchargez contiennent-ils du contenu ?
Si nous entrons un peu plus dans le détail du contenu que vous pouvez attendre des données que vous téléchargez sur GBIF, je vous donne juste quelques idées générales de certains des champs que vous pouvez télécharger.
The GBIF ID, which we apply to the record that is shared with GBIF, is always 100%. You always get all the GBIF IDs for your data. But if you are looking for habitat information, if you look across all occurrence records on GBIF, only around 7% have data in that field.
Individual counts have quite high numbers, as well as state and province, but be mindful that this is because of birds. The biggest group of data we have on GBIF is coming from birds, so they really up how much data we actually have available. So it can be quite group specific.
So if you’re looking for insects, you might find that the percentage of values that are actually covered is quite low.
With scientific name, it’s pretty good.
For the depth, if you’re looking into marine occurrences, very few publishers actually share that data.
The same for elevation, and life stage is only 9%.
Most of our publishers do provide some sort of temporal data, but that could be just in the form of year.
So just be mindful of that when you use your data.

Slide 7 - Data standards
So I’m going to talk a bit about the data standards that are behind what you see on GBIF. As you noticed, the columns have quite specific names. These names and fields are defined by data standards, which are maintained by a global biodiversity community.

Slide 8 - Darwin Core Archive (DwC-A)
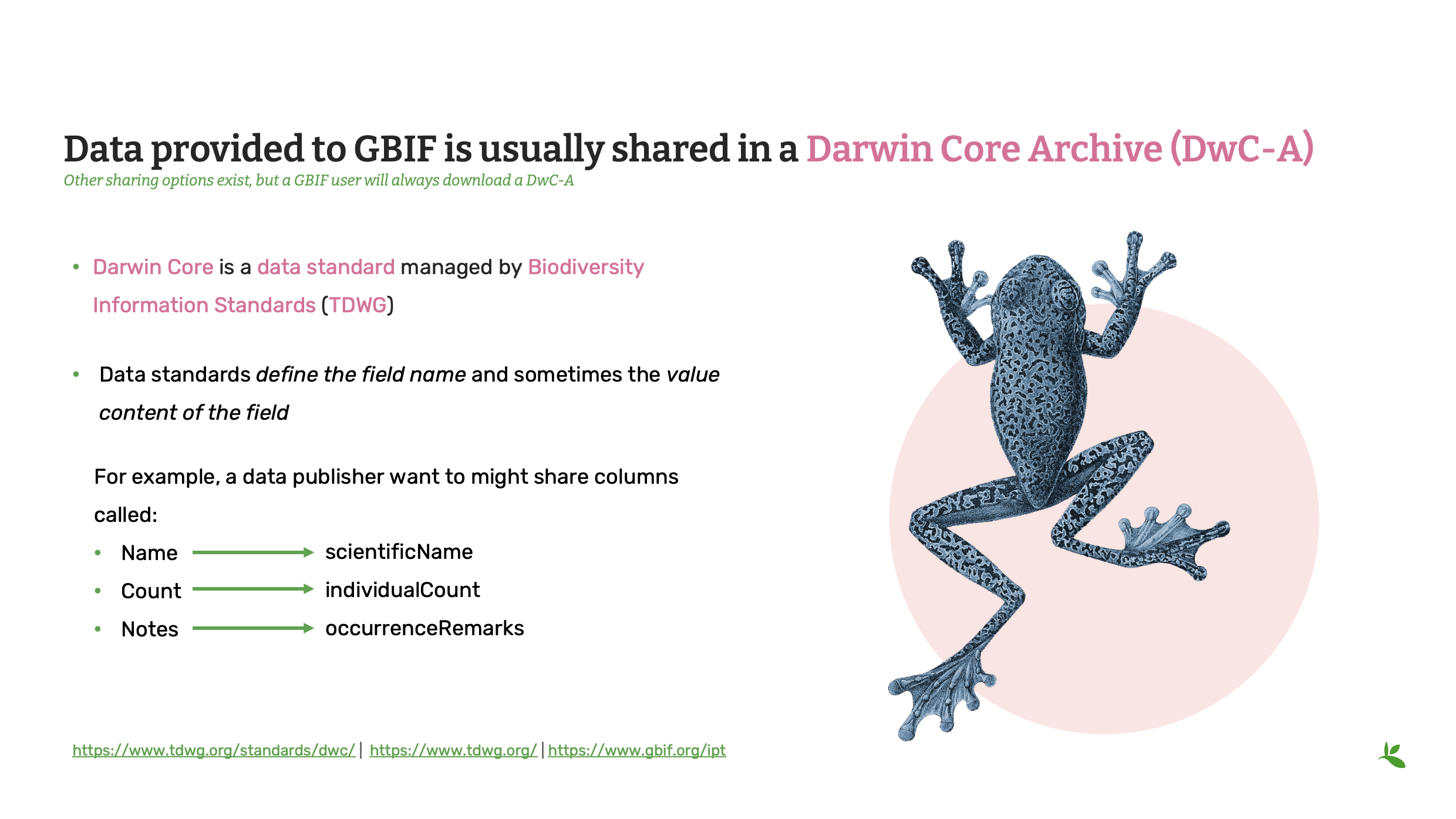
So most of the data shared with GBIF is usually within a Darwin Core archive, as I also mentioned earlier. And the publishers sharing data with GBIF have to map their data to this Darwin Core standard.
It’s maintained by the Biodiversity Information Standards, also called TDWG.
And those data standards define the field name, as I mentioned, and they also sometimes define the value content that could be like a controlled vocabulary or some sort of normalized value that people should use when they share their data.
So let’s say you’re a publisher, you’re completely new to data standards and publishing on GBIF. You might have a spreadsheet saying that you have a name, you have a count of how many of that species you saw, and some general notes about what you found.
Those fields have to be called scientific name for the name, for example, individual count for the counts, and for the notes, you might want to put those in occurrence remarks or event remarks.
The point is that you have specific fields to capture specific information.

Slide 9 - Available fields
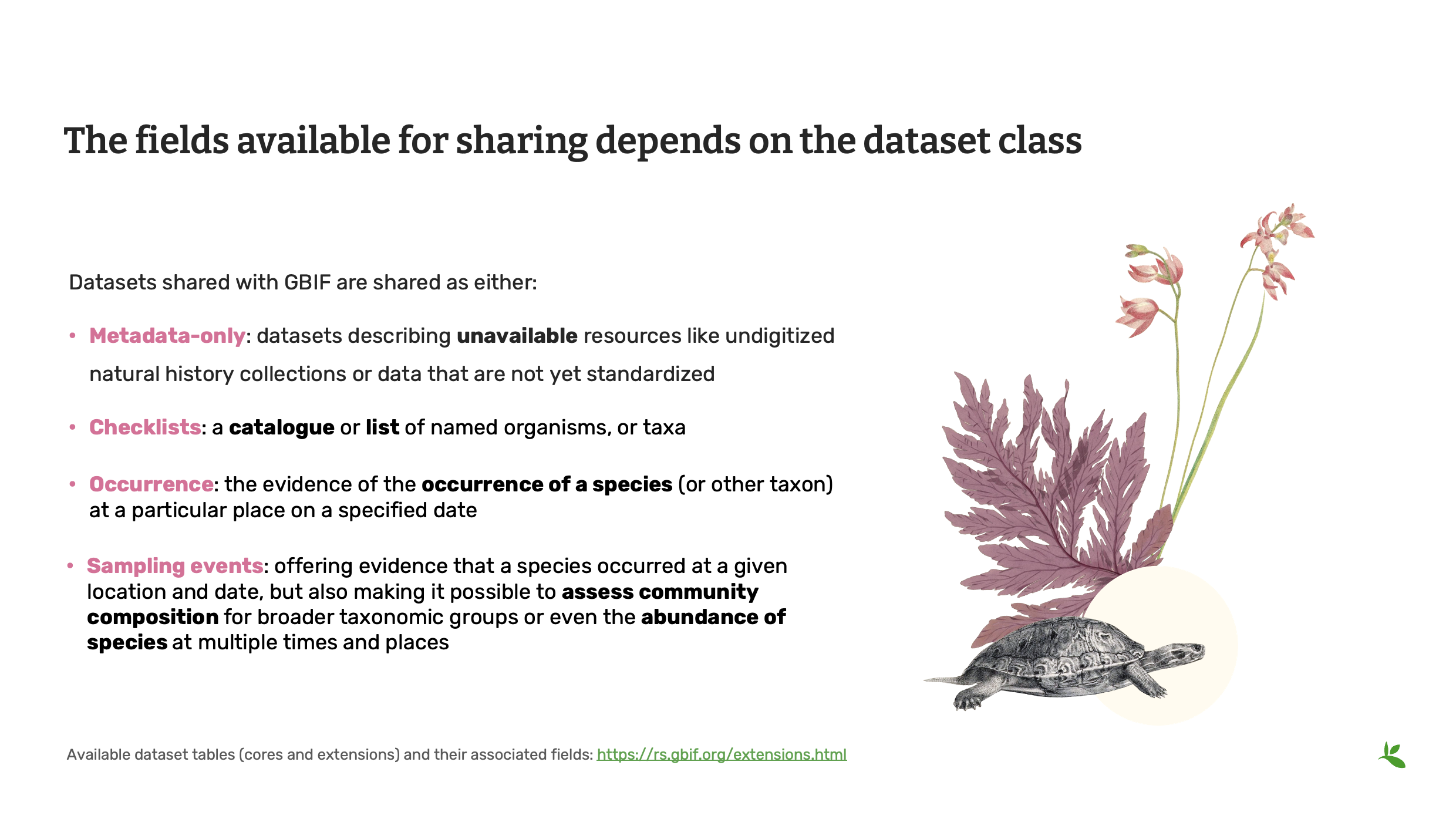
Now, the fields available for sharing on GBIF, it’s dependent on the dataset class.
As I mentioned, you have occurrence records. Those occurrence records is one type of dataset class on GBIF.
So the most simple one is the metadata-only dataset, which doesn’t really have any data. It’s just a description of the dataset. So that could be a description of a collection or description of a field study, but where you haven’t been able to digitize the data just yet.
Then we also have checklist datasets, which is a list of taxon names. There could be more information included, but generally, it’s sharing scientific names of a given area or within a different specific scope.
And then you have occurrences, which I mentioned before, which is the evidence of a species present or a taxon present at a given date and place.
Now, the last type of dataset class we have is the sampling events, which has a little more information because usually people have to share sampling effort and protocol, and it can allow users to assess community composition and abundance of species.
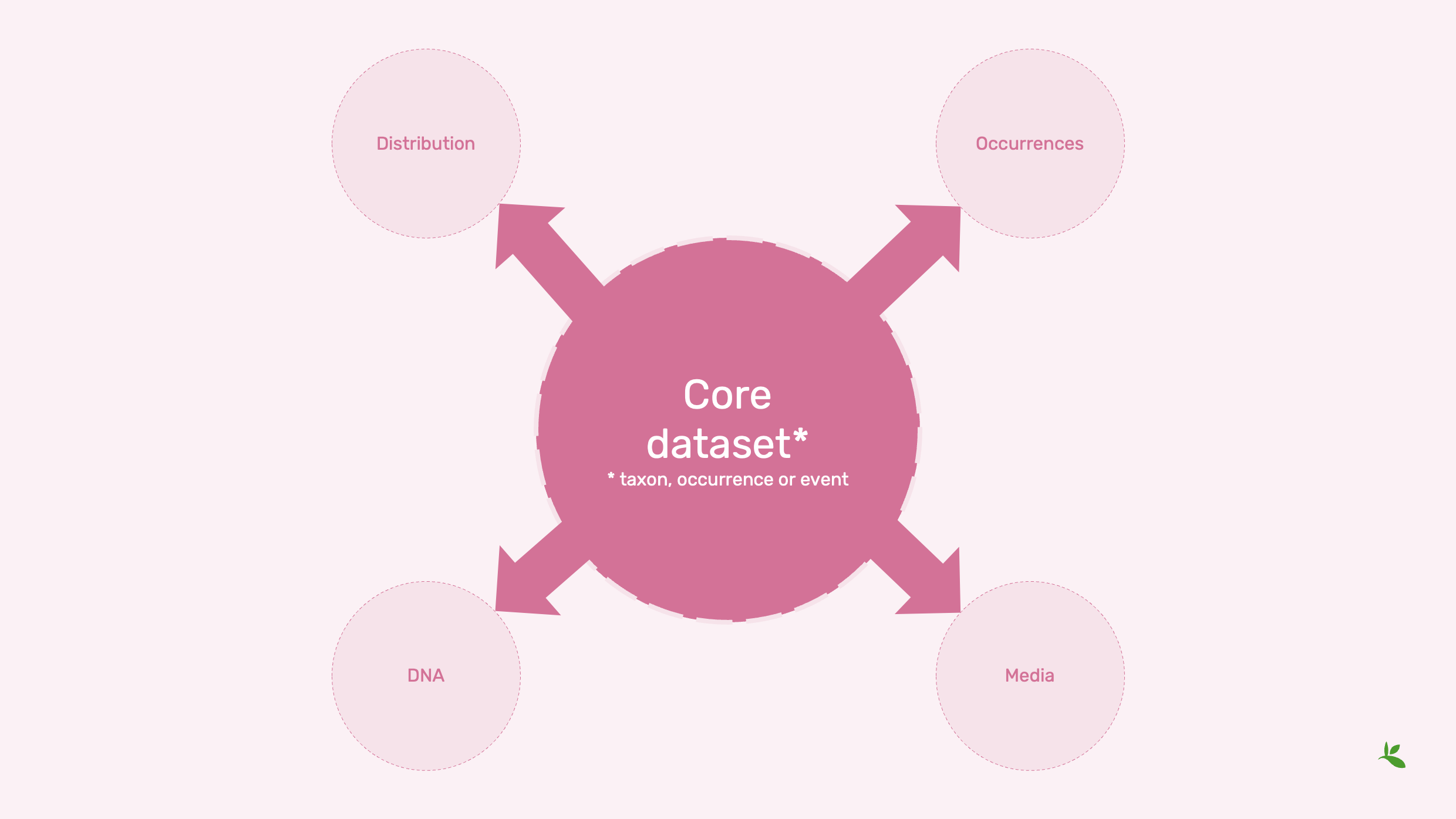
Slide 10 - Dataset cores
Now, those dataset classes are core tables. So if you have a dataset table that is either a taxon, occurrence, or event table those tables have a specific set of fields you can share. But you might want to share even more information.
For example, if you have an event dataset, you might want to share individual occurrences.
You did a BioBlitz from start of Saturday to the end of Saturday, but at that range, you found that butterfly or that beetle at a specific time during that.
That’s an occurrence within the event of the BioBlitz.
Or you might have media associated. You took pictures of the different insects you found during that time, or you might even have taken samples for DNA, and then you have identifications based on DNA.
And you might have a taxon core or a checklist, as I mentioned, where you have the distribution of the species covered.

Slide 11 - Missing fields?
Now, we have a lot of fields available on GBIF in which you can filter down and narrow your search for. But in some cases, you might not find the fields that you need for your analysis.
If you would ever need such a field, the correct way of getting, well, at least starting the discussion of getting that field into GBIF would to be to go through TDWG and the Darwin Core standard.
You can propose new terms and changes to Darwin Core, but you have to be mindful that changing data standards is a community-driven process which requires a lot of engagement and patience because you need to discuss it with your peers, and you need to find a common ground for adding those data. So you have to consider whether the term already exists but would require an update, where the new term would best be placed. Is it taxon specific? Is it occurrence specific or event specific? Is it something to do with locations?
And you also have to think about whether the term is better suited in an extension or in one of the core tables.
So many people are maybe not that aware of extensions to these core tables, but there’s a lot of extensions out there to serve the community who wants to share data or use data.
So please familiarize yourself with them before you propose or change terms.

Slide 12 - Dataset metadata
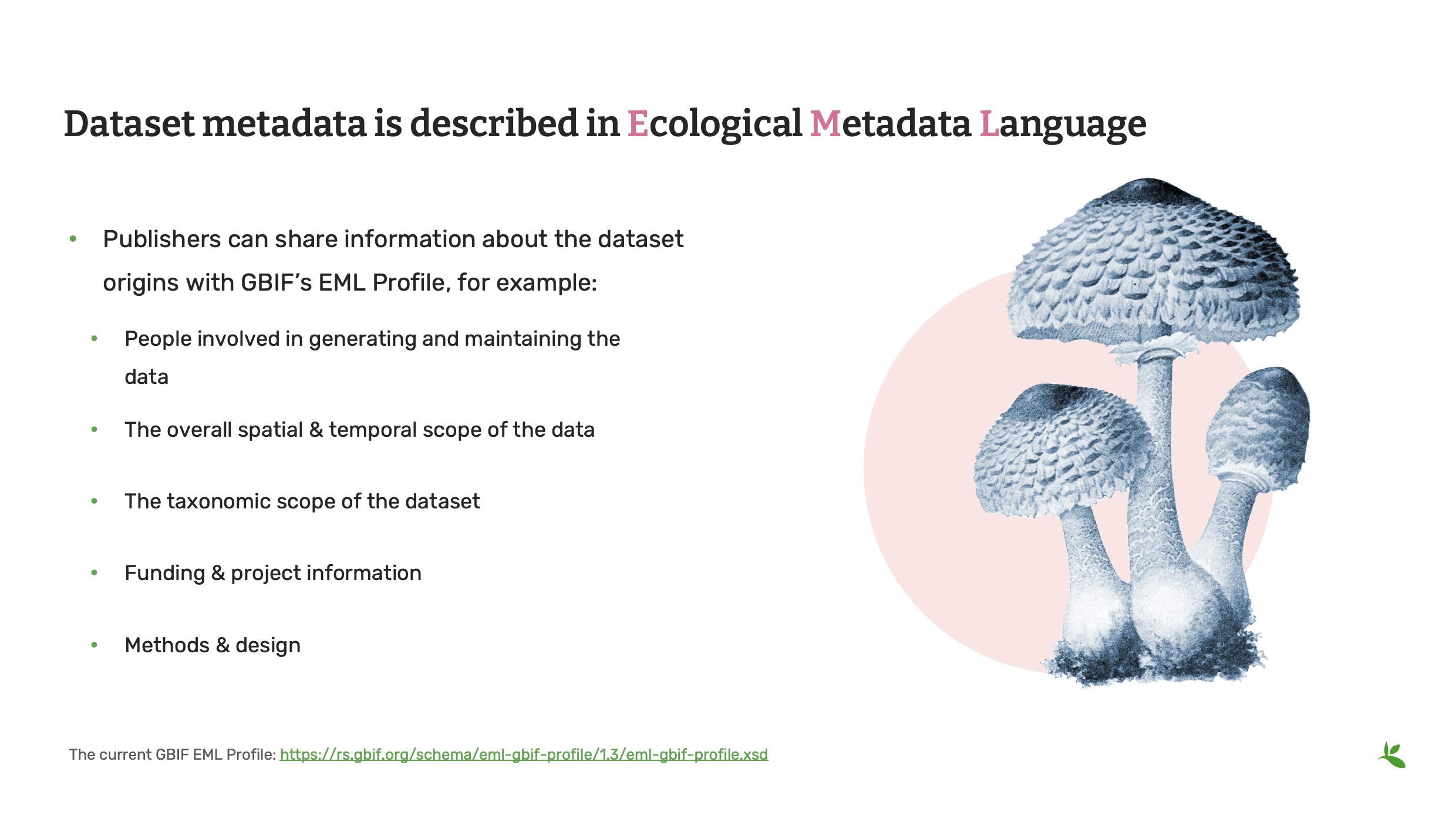
One last thing I have to mention about the data standards is that you also have a data standard called EML or Ecological Metadata Language publishers use to share information about the dataset resource in which they have all their data. So that is, for example, who was the people who was involved in maintaining, gathering the data, the overall spatial and temporal scope of the dataset, the taxonomic scope of the dataset, and any funding and project-specific information, as well as methods and design.
So that’s another standard which is not part of the TDWG standard you might want to look into as well.
Classes de jeux de données GBIF
Plus d’informations sur les classes de jeux de données peuvent être trouvées sur le site Web du GBIF.
Vous pouvez également explorer comment choisir un type de jeu de données.
Interprétation taxonomique
Afin de faciliter la recherche et la génération de mesures, tous les enregistrements d’occurrences sont mis en correspondance avec deux taxonomies :
Catalogue of Life eXtended Release (COL XR) - This is the primary taxonomy used by GBIF.
GBIF backbone - This is a taxonomy that, up until recently, GBIF has been building and integrating periodically. It is primarily based on an older version of Catalogue of Life with additional taxa added in an automated way from other taxonomic datasets. The build process has now been discontinued in favour of the eXtended release of the Catalogue of Life. The GBIF Backbone will no longer be updated, but will remain available for backwards compatibility. All GBIF Backbone identifiers will be preserved and supported in the API.
| API support for taxonomy interpretation along with Taxonomic indexes are documented in GBIF’s technical documentation. |
Why does GBIF need taxonomy interpretation?
Without it, we wouldn’t be able to do any taxonomic search and it would be difficult to generate consistent statistics and maps.
As you can imagine, not everyone uses the same classifications or names. This results in considerable variations in higher taxa and a large number of synonyms. This process aims to bring all these names together and organize them.
How are the taxonomies generated?
Catalogue of Life is a global index of species, which aims to provide a comprehensive and authoritative list of the world’s species. It is compiled from multiple taxonomic datasets and is updated regularly. The eXtended Release (COL XR) builds on the Base Release by programmatically integrating additional data sources. It integrates information from over 58,000 overlapping taxonomic and nomenclatural global, regional, national and management data sources (checklists) as well as originating from digitised literature available in Catalogue of Life’s infrastructure ChecklistBank.
| Notez que de nombreuses occurrences basées sur des séquences génétiques n’ont pas de noms latins, mais sont nommées en utilisant des hypothèses d’espèces (UNITE: champignons) ou des Indices Numériques de Codes-Barres (iBOL : principalement des animaux). C’est pourquoi l’ajout de ces deux principales sources d’OTUs à la dernière version de l’ossature taxonomique améliore considérablement la fonctionnalité d’indexation du GBIF pour les données de biodiversité basées sur des séquences ADN. |
How are the taxonimic status' defined?
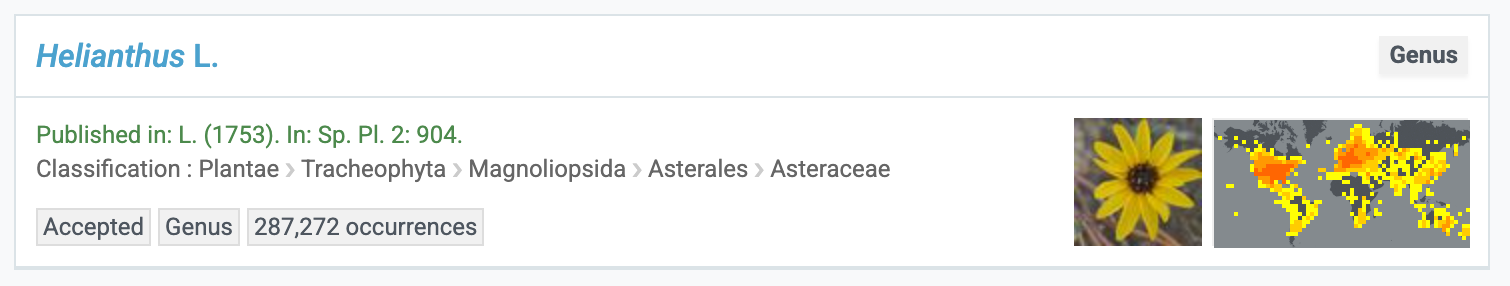
-
Accepted: A taxonomically accepted, current name
-
Provisionally accepted: Treated as accepted, but doubtful whether this is correct.
-
Synonym: Names which point unambiguously at one species (not specifying whether homo- or heterotypic).Synonyms, in the CoL sense, include also orthographic variants and published misspellings.
-
Ambiguous synonym: Names which are ambiguous because they point at the current species and one or more others e.g. homonyms, pro-parte synonyms (in other words, names which appear more than in one place in the Catalogue).
-
Misapplied: A misapplied name. Usually accompanied with an accordingTo on the synonym to indicate the source the misapplication can be found in.
-
Bare name: A name alone without any usage, neither a synonym nor a taxon.
GRSciColl
The Global Registry of Scientific Collections, or GRSciColl, is a comprehensive and community-curated clearing house of information about scientific collections in the GBIF registry. By providing information about physical scientific collections—their content, location, contacts, associated institutions, and collection codes and identifiers—GRSciColl offers a resource for a wide range of uses by experts, researchers and members of broader society.
|
In this video (02:11), Marie Grosjean, GBIF Data Administrator, provides an introduction to GRSciColl. If you are unable to watch the embedded Vimeo video, you can download it locally (MP4 - 13.3 MB). |
Principles of GBIF-mediated data
| In this section, you will learn about the principles that GBIF follows with regards to data and how data in the GBIF portal are FAIR. |
Digital object identifiers
A Digital Object Identifier, or DOI, is a standard, permanent identifier that provides an actionable, interoperable, persistent link to any entity. The concept is that DOI differs from commonly used references like URL web links because it identifies an object itself as a first-class entity, not simply the place where the object is currently located.
In the context of GBIF.org, DOIs serve as stable identifiers for four different types of things:
-
datasets from the GBIF network
-
data downloads from GBIF.org
-
research articles and reports published by scientific journals, agencies and NGOs
-
materials deposited in a general-use repository
Le GBIF assigne des DOI à tous les jeux de données et téléchargements d’occurrences. Lorsque des données sont utilisées, suivre les règles de citation DOI offre une façon facile et cohérente de créditer les détenteurs de jeux de données tout en permettant la reproductibilité. Les DOI se rapporteront toujours au jeu de données ou aux pages de téléchargement, même si les données sous-jacentes ne sont plus disponibles.
GBIF started issuing DOIs on 3 February 2015. Downloads requested before this date do not have DOIs, however, if you wish to cite older downloads, you can contact helpdesk@gbif.org and we will assign DOIs as appropriate.
Standards
The data available through GBIF.org and its associated services is the result of the GBIF network of Participants and publishers applying shared rules and conventions to describe, record and structure thousands of different datasets drawn from hundreds of institutions around the world. Common standards are the main enabler for bringing together the hundreds of millions of primary biodiversity records in the GBIF index.
Within the biodiversity domain, the group most often responsible for developing and maintaining data standards is Biodiversity Information Standards. This nonprofit scientific and educational association focuses on the development of standards for the exchange of biological and biodiversity data. Members of the biodiversity community generally refer to this group as TDWG (pronounced tad-wig)—a vestigial reminder of its earlier manifestation as the Taxonomic Databases Working Group.
Les standards utilisés fréquemment comprennent :
-
Darwin Core: The Darwin Core Standard (DwC) offers a stable, straightforward and flexible framework for compiling biodiversity data from varied and variable sources. The majority of the datasets shared through GBIF.org are published using the Darwin Core Archive format (DwC-A).
-
Ecological Metadata Language (EML): Ecological Metadata Language is a metadata standard that records information about ecological datasets in a series of modular and extensible XML document types. All of the descriptions of datasets in GBIF.org rely on ‘metadata’—that is, the information about data—using the open-source EML standard, which is administered and maintained by The Knowledge Network for Biocomplexity. Each Darwin Core Archive includes as one of its components an EML file (written in XML format).
-
BioCASe/ABCD: The Biological Collection Access Service, commonly referred to as BioCASe, is an international network linking biological collections data from natural history museums, botanical/zoological gardens and research institutions. The BioCASe protocol relies on the Access to Biological Collections Data (ABCD) data exchange standard, which TDWG also administers.
Open data
In keeping with a 2014 decision by the GBIF governing board, data publishers must assign one of the three Creative Commons options to any occurrence dataset. The Governing Board recognized the need for much greater clarity both for data publishers and users on how data may be used when shared via GBIF.org. Creative Commons is a nonprofit organization that helps overcome legal obstacles to the sharing of knowledge and creativity to address the world’s pressing challenges.
| Notez que la licence CC-BY-NC a un effet significatif sur la possibilité de réutiliser les données. Le GBIF encourage les fournisseurs de données à choisir l’option la plus ouverte possible. Il est important de noter que les images ne sont pas soumises à la même licence qui est appliquée au jeu de données et peuvent avoir des conditions d’utilisation plus restreintes. Enfin, l’attribution/citation est une norme communautaire, donc même si les éditeurs ont renoncé à des conditions d’utilisation, l’attribution est attendue. |
FAIR data
Many articles from 2011-2016 documented a crisis in scientific reproducibility (see below). In 2016, the FAIR Guiding Principles for scientific data management and stewardship were published in Scientific Data. The principles were designed to improve the Findability, Accessibility, the Interoperability and the Reusability of datasets and address "an urgent need to improve the infrastructure supporting the reuse of scholarly data." Implementation of these principles began in 2018. You can read more about How to GO FAIR on GO-FAIR.org.
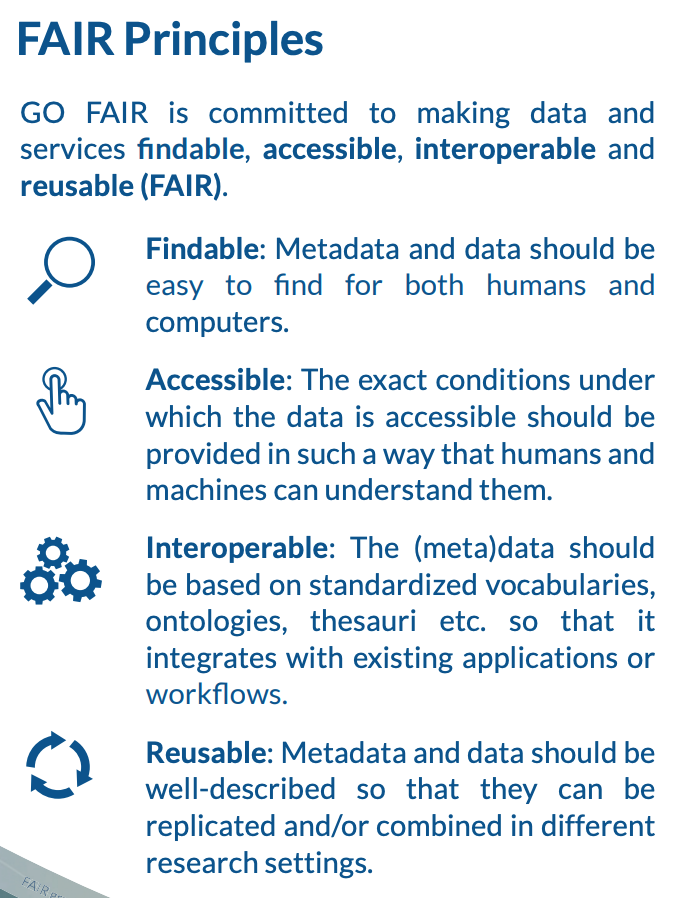

Data found on GBIF.org are FAIR.
Literature references
Baker (2016) 1,500 scientists lift the lid on reproducibility. Nature 533: 452-454 (26 May 2016) doi:10.1038/533452a
Baker (2016) Reproducibility: Seek out stronger science. Nature 537: 703-704 (29 September 2016) doi:10.1038/nj7622-703a
Nature editorial (2016) Reality check on reproducibility. Nature 533: 437 (26 May 2016) doi:10.1038/533437a
Baker (2016) Statisticians issue warning over misuse of P values. Nature 531: 151 (10 March 2016) doi:10.1038/nature.2016.19503
Nosek et al. (2015) Promoting an open research culture. Science 348(6242): 1422-1425. DOI:10.1126/science.aab2374
Leek and Peng (2015) Statistics: P values are just the tip of the iceberg. Nature 520: 612 (30 April 2015) doi:10.1038/520612°
Nuzzo (2015) How scientists fool themselves – and how they can stop. Nature 526: 182–185 (08 October 2015) doi:10.1038/526182a
Hayden (2013) Weak statistical standards implicated in scientific irreproducibility. Nature doi:10.1038/nature.2013.14131
Young (2012) Replication studies: Bad copy. Nature 485, 298–300 (17 May 2012) doi:10.1038/485298a
Callaway (2011) Reports finds massive fraud at Dutch universities. Nature 479, 15 (1 November 2011) doi:10.1038/479015a
Statistiques des données
| Dans cette section, nous passons en revue les différentes statistiques disponibles pour les ensembles de données. |
L’un des nombreux avantages de la publication de données via le GBIF est que, durant le processus d’indexation, le GBIF analyse tous les jeux de données et produit des statistiques à leur sujet. Ces statistiques sont mises à disposition de plusieurs manières :
-
tendances globales
-
pages des pays
-
statistiques sur le contenu du jeu de données
-
activité de téléchargement du jeu de données
Participants and publishers can use this information to improve the quality of their datasets, e.g. by addressing issues detected during the indexing process. They can also use the access statistics as evidence of real user interest in their datasets and potential use of the published data.
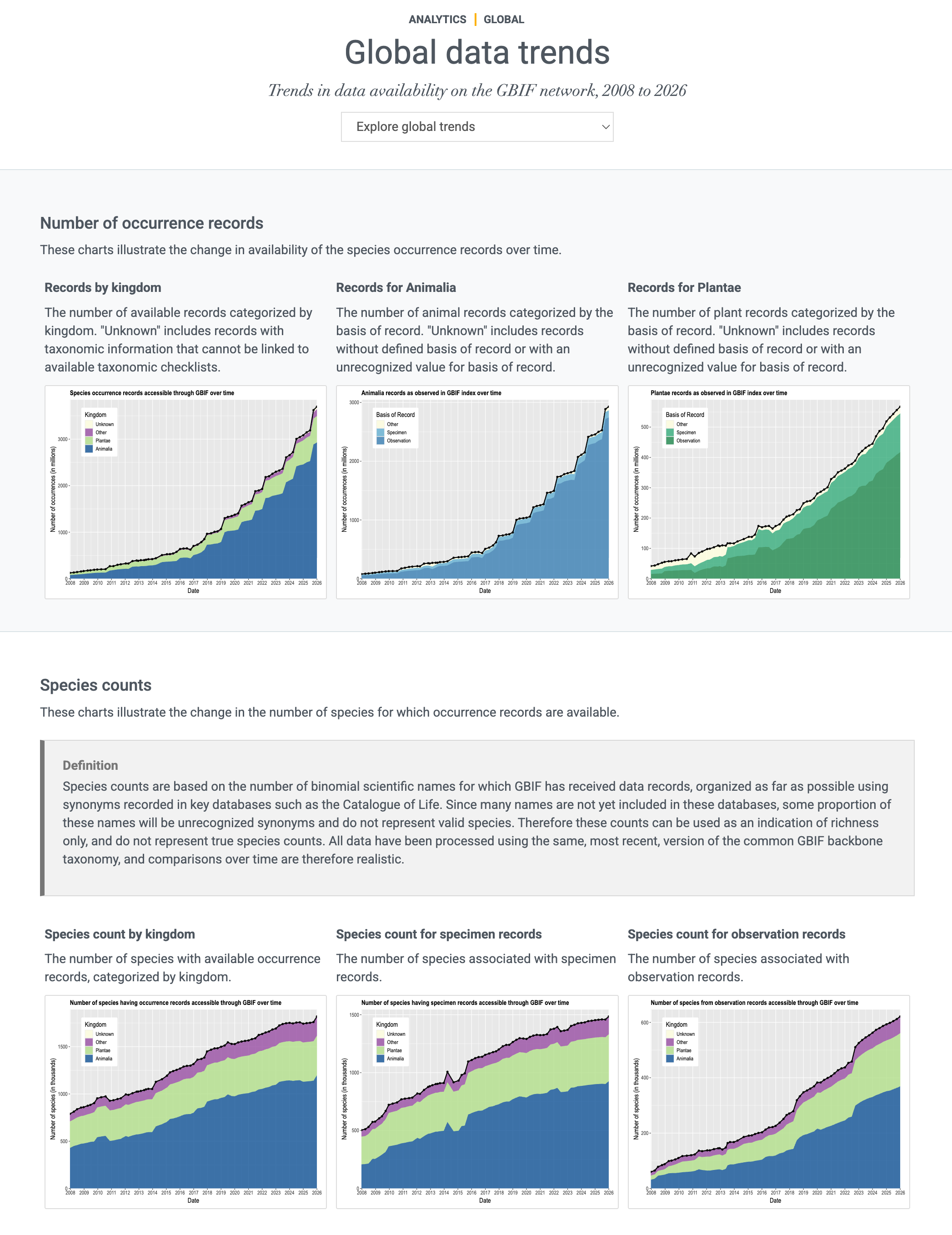
Global data trends
GBIF.org met régulièrement à jour les statistiques pour fournir un aperçu des tendances globales des données de 2008 à nos jours. Les graphiques illustrent les tendances dans :
-
relevés d’occurrences
-
nombre d’espèces
-
temps et saisonnalité
-
exhaustivité et précision
-
couverture géographique des espèces recensées
-
partage de données avec le pays d’origine
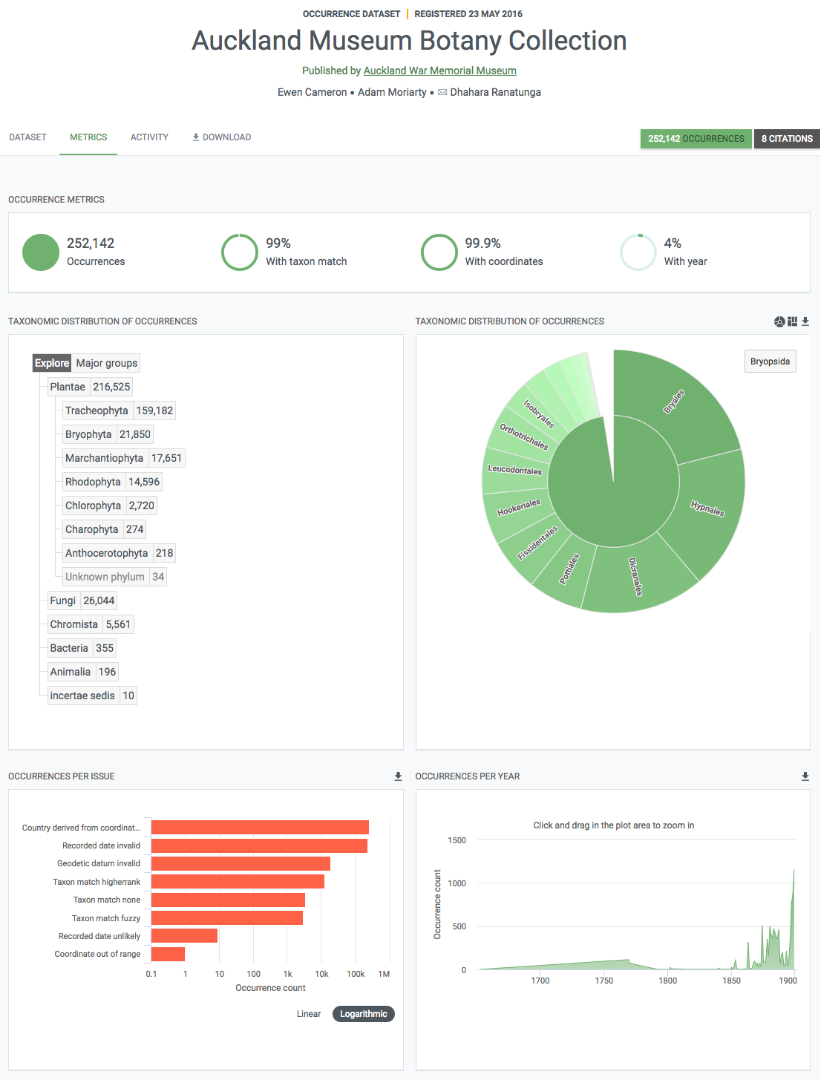
Statistiques sur le contenu du jeu de données
Chaque page du jeu de données comprend un onglet intitulé "Statistiques". Cet onglet donne accès aux graphiques et aux tableaux résultant de l’analyse du contenu du jeu de données. Cela inclut des résumés de :
-
La distribution taxonomique (sous forme de liste et de graphique)
-
Occurrences par problème
-
Occurrences par année
Les graphiques/tableaux sont interactifs et vous pouvez cliquer dessus pour filtrer et explorer. De plus, les images peuvent être téléchargées à des fins de rapports.
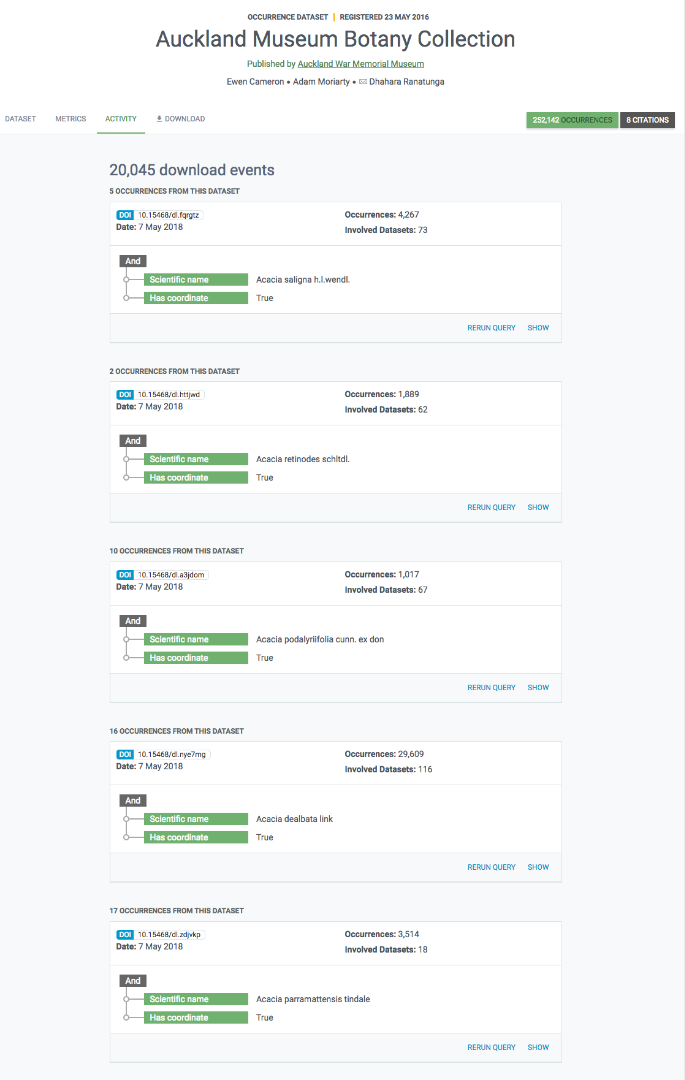
Les rapports d’accès aux données
Il y a un troisième onglet dans les pages du jeu de données des occurrences intitulé « Activité ». Dans cet onglet, vous pouvez voir une liste de toutes les demandes de téléchargement qui ont inclus des enregistrements de ce jeu de données, y compris leur DOI de téléchargement pour un suivi facile.
Révision
|
Testez vos connaissances sur les concepts abordés dans ce module. Certaines questions peuvent comporter plusieurs réponses correctes. Vous pouvez en savoir plus sur les réponses dans le Annexe des solutions. |