Publication de données
|
Dans ce module, vous apprendrez des concepts sur la publication de données, incluant l’IPT, les cœurs (cores) et les extensions, et l’importance des licences, des métadonnées, des champs obligatoires et de l’hébergement des jeux de données. |
Concepts de publication de données
|
In this video (15:27), you will learn about data publishing concepts and will receive an introduction to the Integrated Publishing Toolkit (IPT). Subtitles are not yet available for this video. If you are unable to watch the embedded video, you can download it locally. (MP4 - 53.2 MB) If you prefer to read, you will find the transcript below the embedded video. |
Transcription de la présentation
Cliquez pour développer

Slide 1 - Data publishing concepts
In this presentation, we will introduce you to the concept of Data Publishing. We will also introduce you to GBIF’s Integrated Publishing Toolkit.
Photo: Euphorbia royleana Boiss. Observed in Nepal by Yaling Lin CC BY-NC 4.0
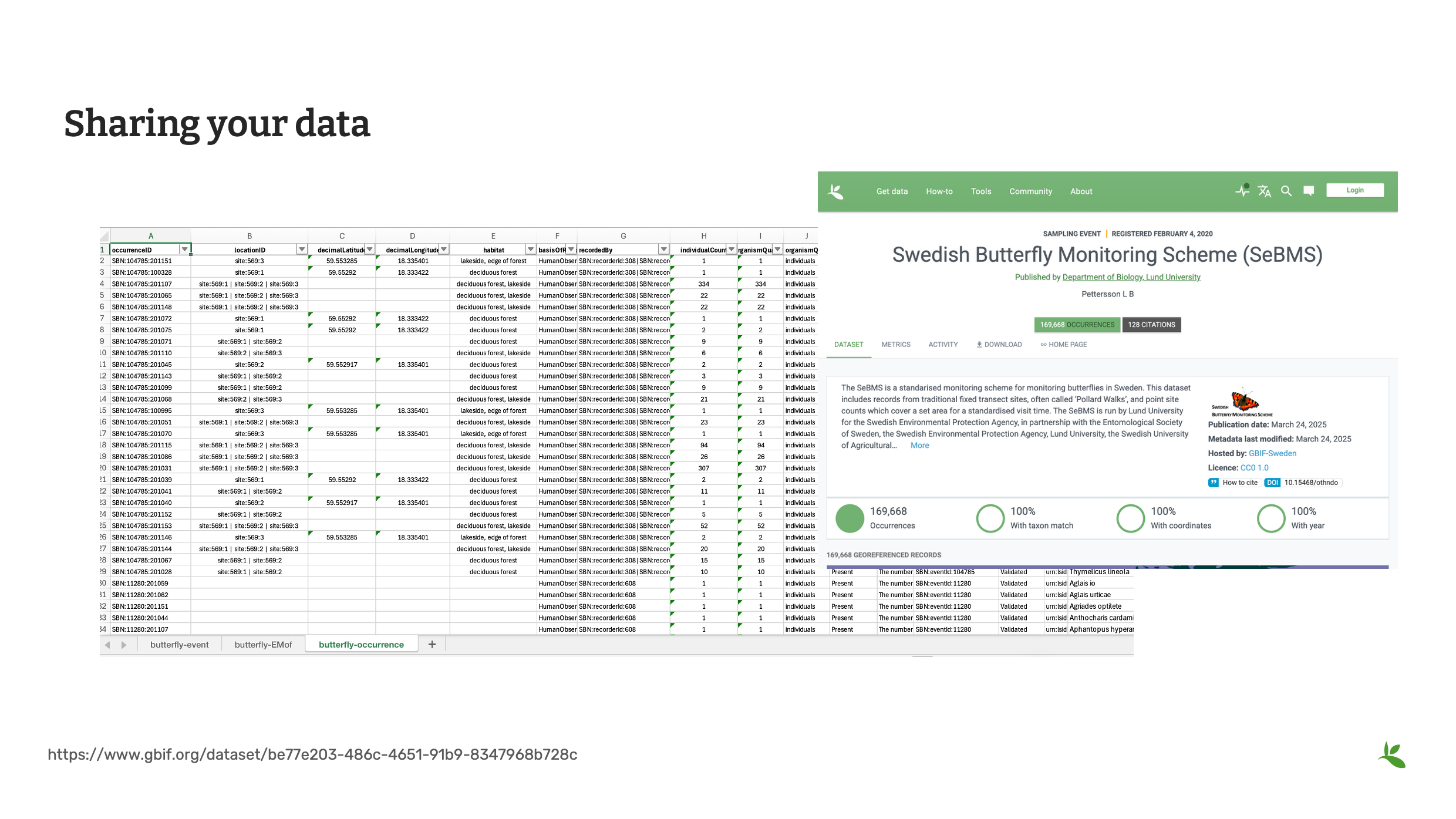

Slide 2 - Sharing your data
At this point in the course, you have moved through all the sections of a mobilization project. Now you are ready to share your data. You’ve produced a cleaned and standardized dataset and you want to share and see that data on the GBIF data portal.
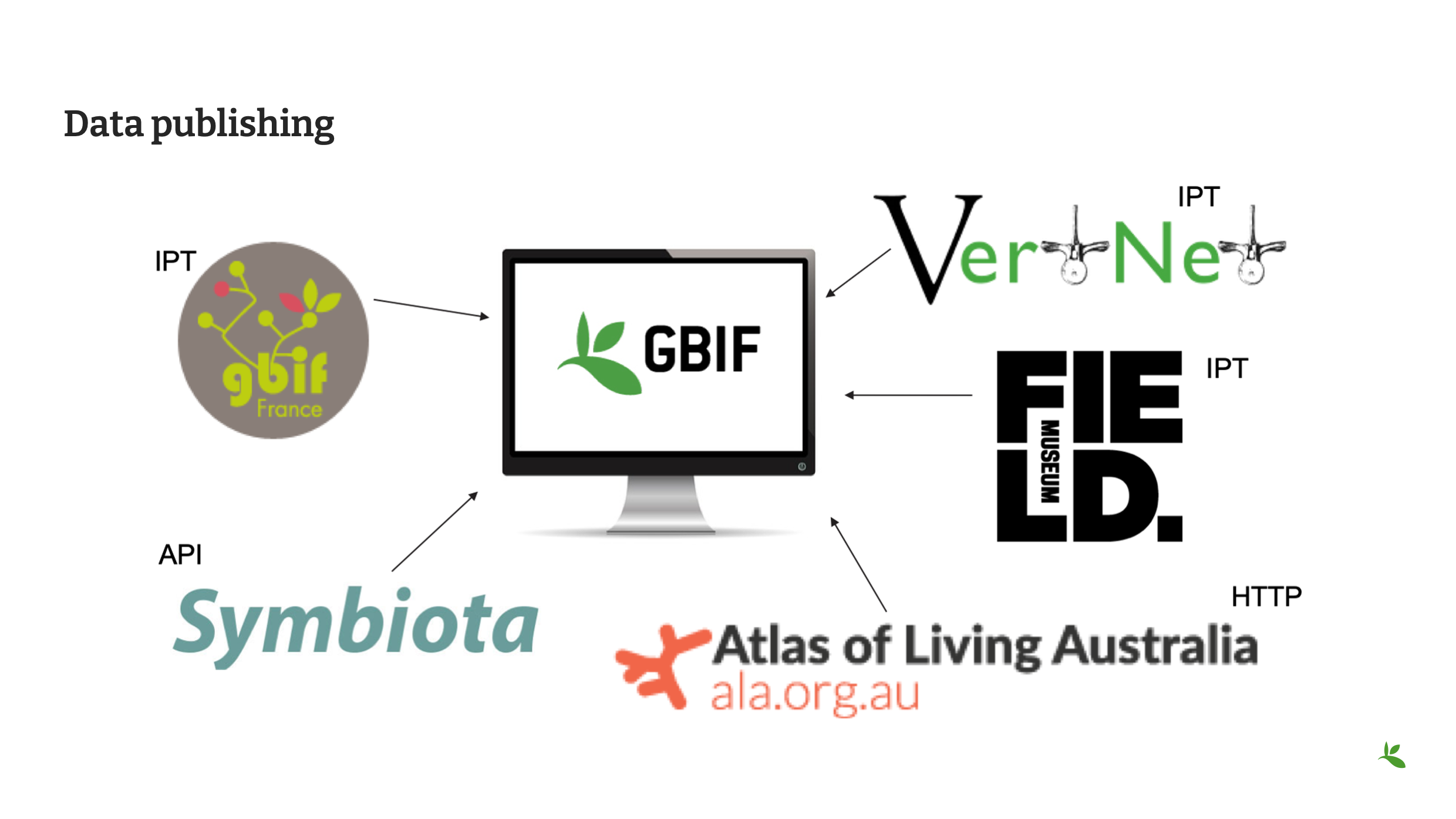
Slide 3 - Data publishing
But there is more to it than simply handing over a spreadsheet or a text file to GBIF. You must publish the data. Data publishing is the act of making biodiversity datasets publicly accessible, discoverable and as open as possible, in a standardized form (i.e. Darwin Core), via an access point. This access point is a URL – a web address. Most organizations within the GBIF Network, otherwise known as publishers, make use of an IPT, the Integrated Publishing Toolkit, to publish their data. These publishers may choose to host their own installation of an IPT like the Field Museum, but generally they prefer to find a suitable host for their data publishing activities. This might be through an established GBIF Participant Node or through an established thematic group like VertNet. Or, an another option, might be to make use of one of the GBIF regional IPTs. If you are part of groups like Symbiota or the Living Atlases Communities, they have other means to assist you with publishing your data to GBIF.

Slide 4 - Creative Commons waiver and licenses
The previous slide mentions the act of not only making the data accessible, but making it as open as possible.
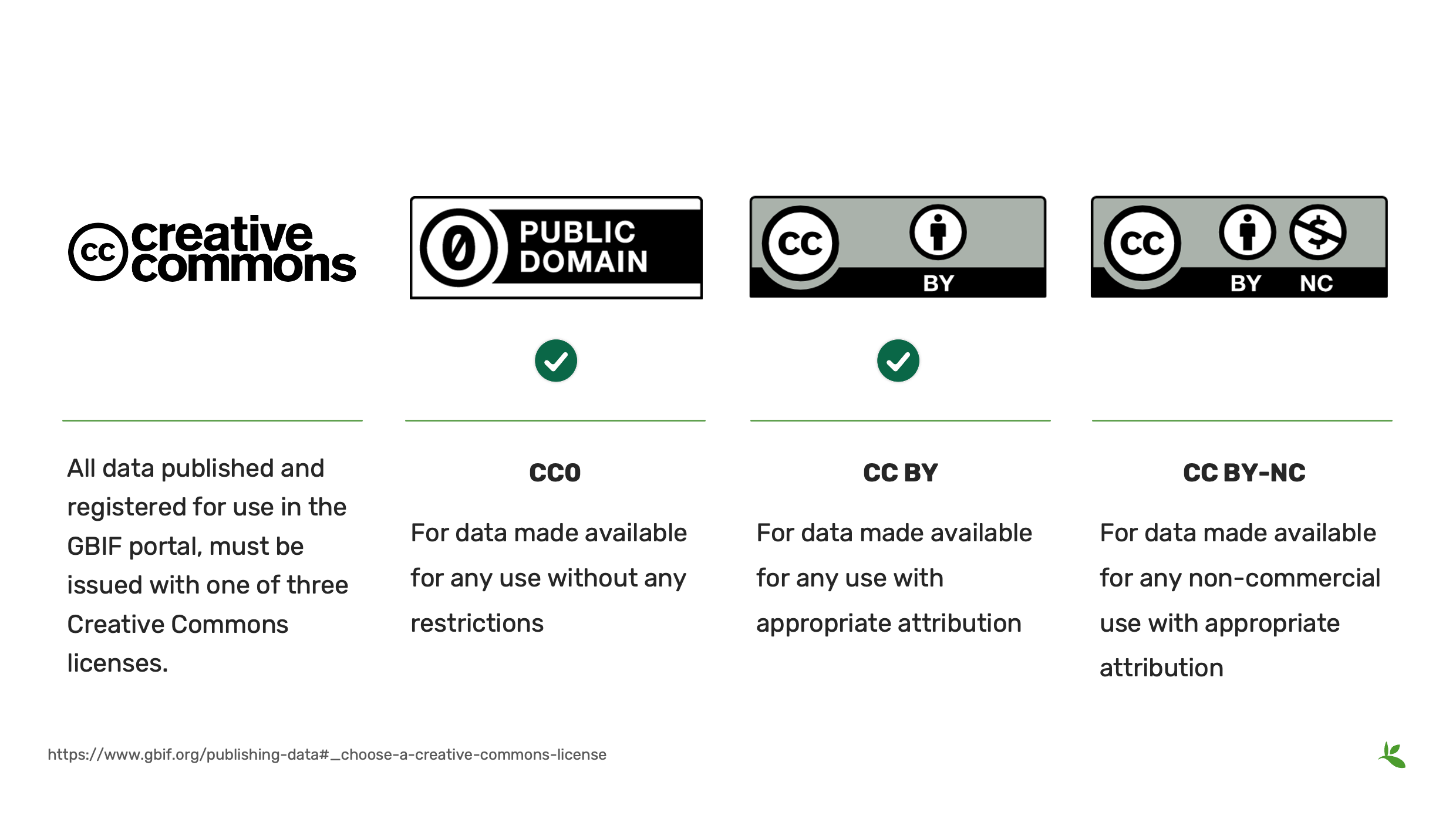
In an effort to document the level of openness of a dataset, GBIF maintains, in keeping with a 2014 decision by the GBIF governing board, that all data published and registered for use in the GBIF portal, must be issued with one of three Creative Commons options:
-
CC0, a waiver for data made available for any use without any restrictions
-
CC BY, a license for data made available for any use with appropriate attribution
-
CC BY-NC, a license for data made available for any non-commercial use with appropriate attribution.
INFORMATION: CC-BY-NC licences have a significant effect on the reusability of data.
GBIF encourages data publishers to choose the most open option they can wherever possible.
Projects supported through GBIF-led funding must use either the CC0 or CC BY options.

Slide 5 - Integrated Publishing Toolkit
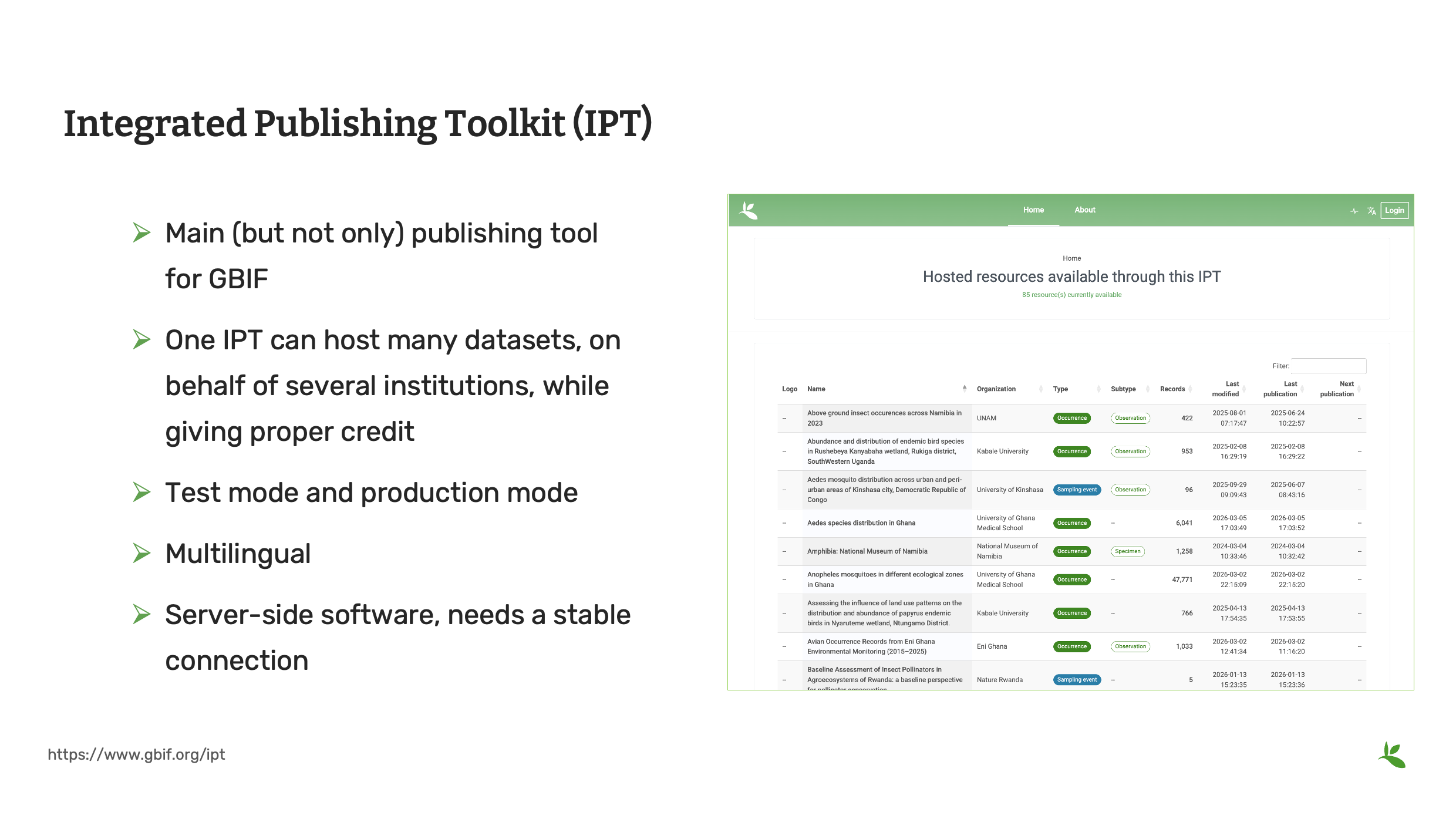
The Integrated Publishing Toolkit (IPT) is a free, open-source software tool written in Java that is used to publish and share biodiversity datasets through the GBIF network. The first version of IPT was released in 2009. It was redesigned and rereleased in 2011. Since that time, the IPT has been updated continuously updated with new features, bug fixes and security improvements.
While the IPT is maintained at GBIF, the community of biodiversity informatics developers are able to contribute to its development.
Some of the features include:
-
Main (but not only) publishing tool for GBIF.
-
One IPT can host many datasets, on behalf of several institutions, each clearly represented
-
Test mode and production mode
-
Multilingual – it has been translated into 7 languages.
It is a server-side software, so it needs a stable connection and requires technical administration. This is why many institutions choose to publish their data on hosted IPTs.
You can find more information on data hosting on the GBIF website.
For the remainder of our course, we’ll be demonstrating and completing exercises using the IPT.

Slide 6 - Choosing a publishing core
We’re now putting together what you learned in the first two presentations in the Data Capture section on Standards and Darwin Core and Data Types. At its heart, the IPT is based on Darwin Core and the Darwin Core text guide and its recommendations for sharing data in a Darwin Core Archive. The Darwin Core contains three distinct data cores which match up to three of the dataset types that GBIF accepts. Keep in mind: * Each dataset has one and only one Core file * Each Core file corresponds to a dataset type: occurrence, checklist or sampling event. * Each dataset can have one or more extensions file * The aim of an extension is to add new data fields not present in the core. * Each entry in the core file is linked to zero, one or several rows in an extension file. * Each row in an extension file reference one and only one row in the core file.
Some examples:
-
Core is Occurrence, no extensions
-
Core is Occurrence, with “multimedia” extension
-
Core is Occurrence, with simple multimedia and measurementorfact extensions
-
Core is Taxon, no extensions
-
Core is Taxon, with species distribution extension
-
Core is Event with occurrence and measurementorfact extensions
Photo: Pieris formosa (Wall.) D.Don Observed in Nepal by Neil Alderson http://creativecommons.org/publicdomain/zero/1.0/[CC0 1.0]
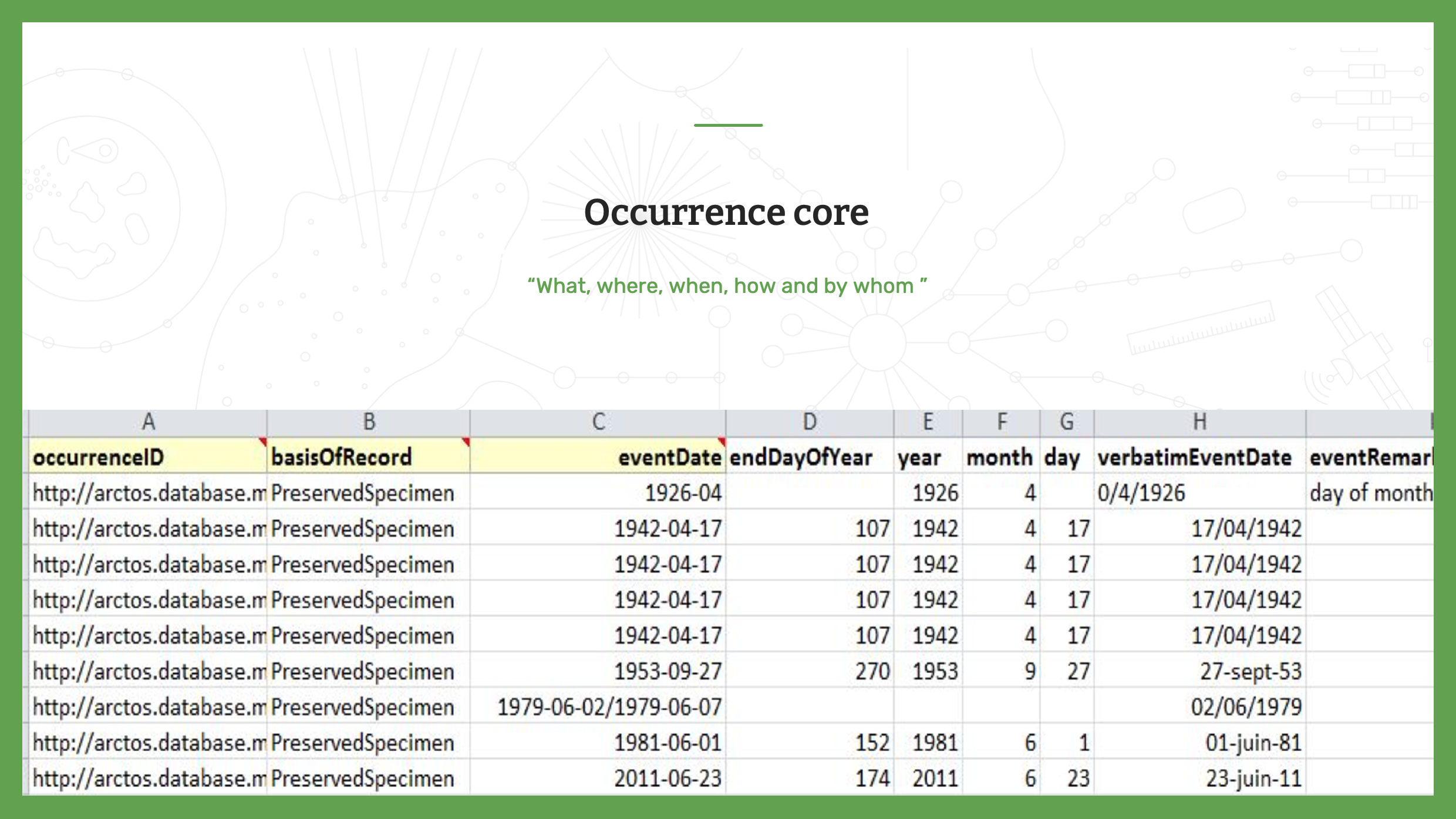
Slide 7 - Occurrence Core
The first core is the Occurrence Core. As a reminder, an occurrence dataset contains one individual or one group of individuals. Each row has a unique identifier, an occurrenceID. Other fields for occurrence data include where, when, how and by whom was each occurrence observed and/or collected in the field.

Slide 8 - Taxon Core
The next option is the Taxon core. As a reminder, a checklist dataset contains taxonomic concepts, not individuals. It is a “A catalogue or list of named organisms, or taxa.” Each row will have a taxonID. Each taxon must be unique.
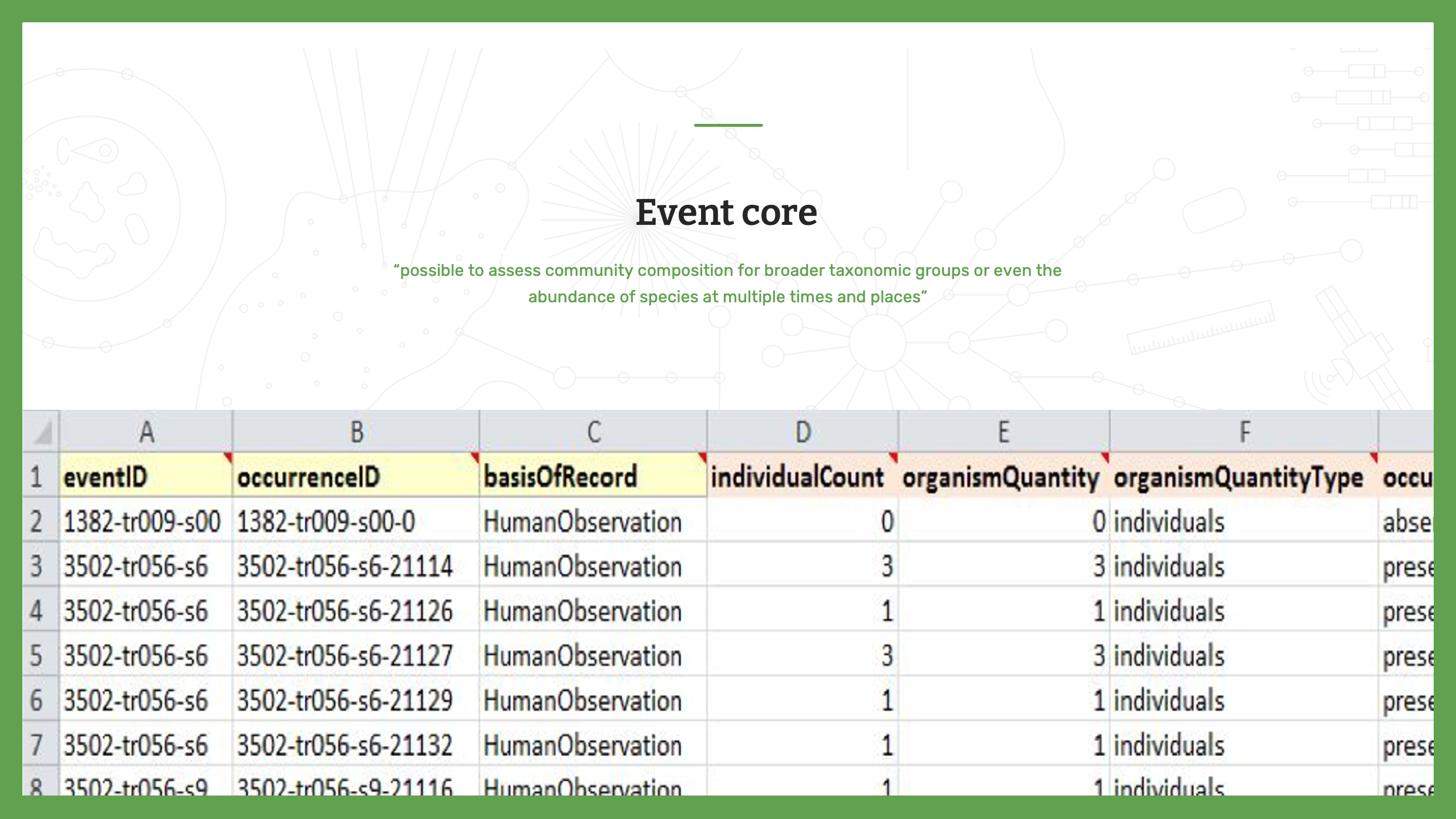
Slide 9 - Event Core
The last option is the Event core. As a reminder, sampling event datasets allow data publishers to provide greater detail, not only offering evidence that a species occurred at a given location and date, but also making it possible to assess community composition for broader taxonomic groups or even the abundance of species at multiple times and places. By indicating the methods, events and relative abundance of species recorded in a sample, these datasets improve comparisons with data collected using the same protocols at different times and places—in some cases, even leading researchers to infer the absence of particular species from particular sites.
Each row will have an EventID. Each event must be unique.
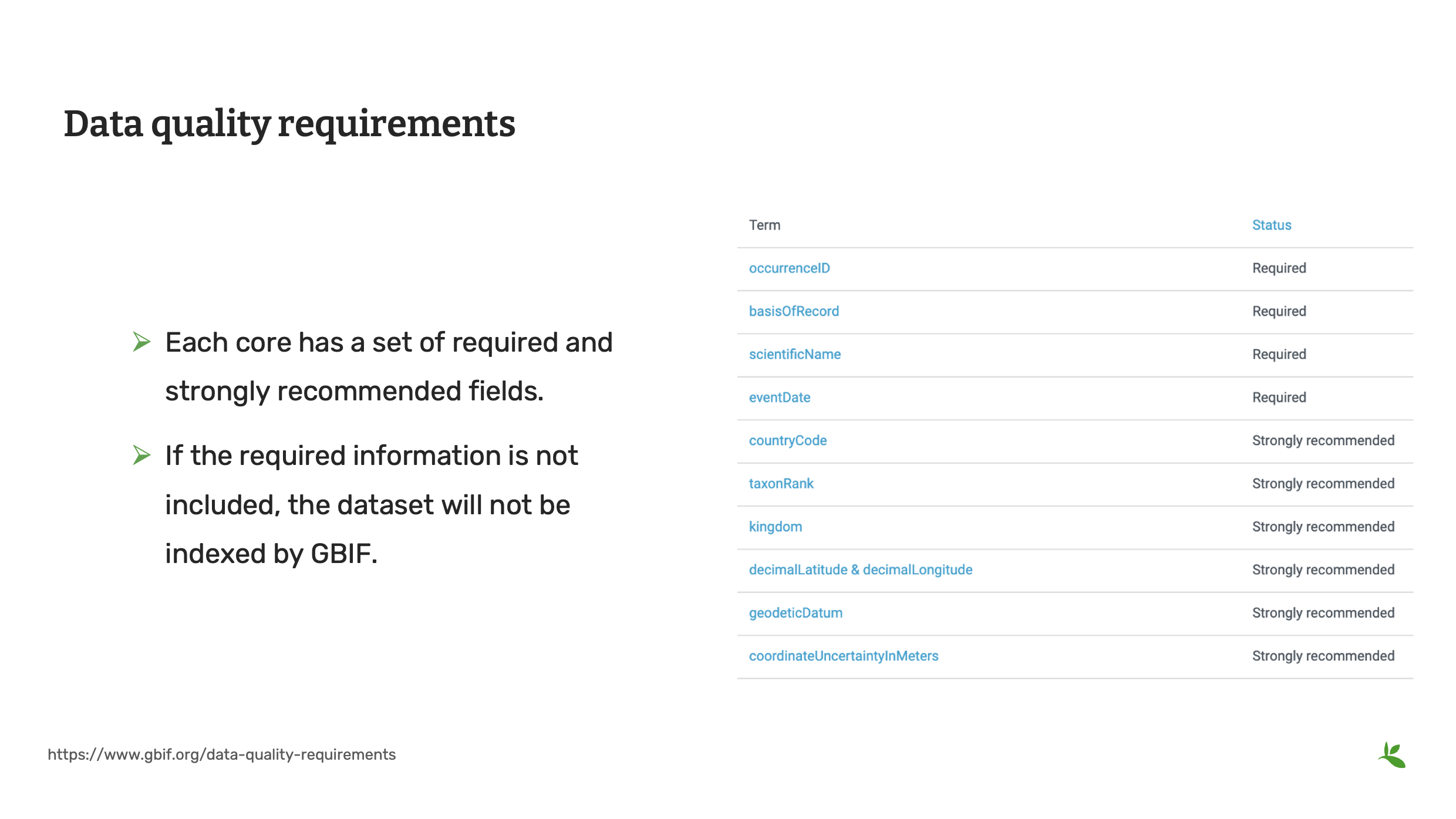
Slide 10 - Data quality requirement
This is a good point to review all of the fields required by GBIF. If required fields are not supplied, GBIF will have difficult indexing your dataset.
Each core has its own set of required and recommended fields for the dataset and for the associated dataset metadata. As you now know, metadata is the data about your data and it enables users to know if your dataset will be fit for their use.
Don’t stop at the required and recommended fields. The more data you are able to share, the more useful your dataset can be to end users.
You can find the data quality requirments on the GBIF website.
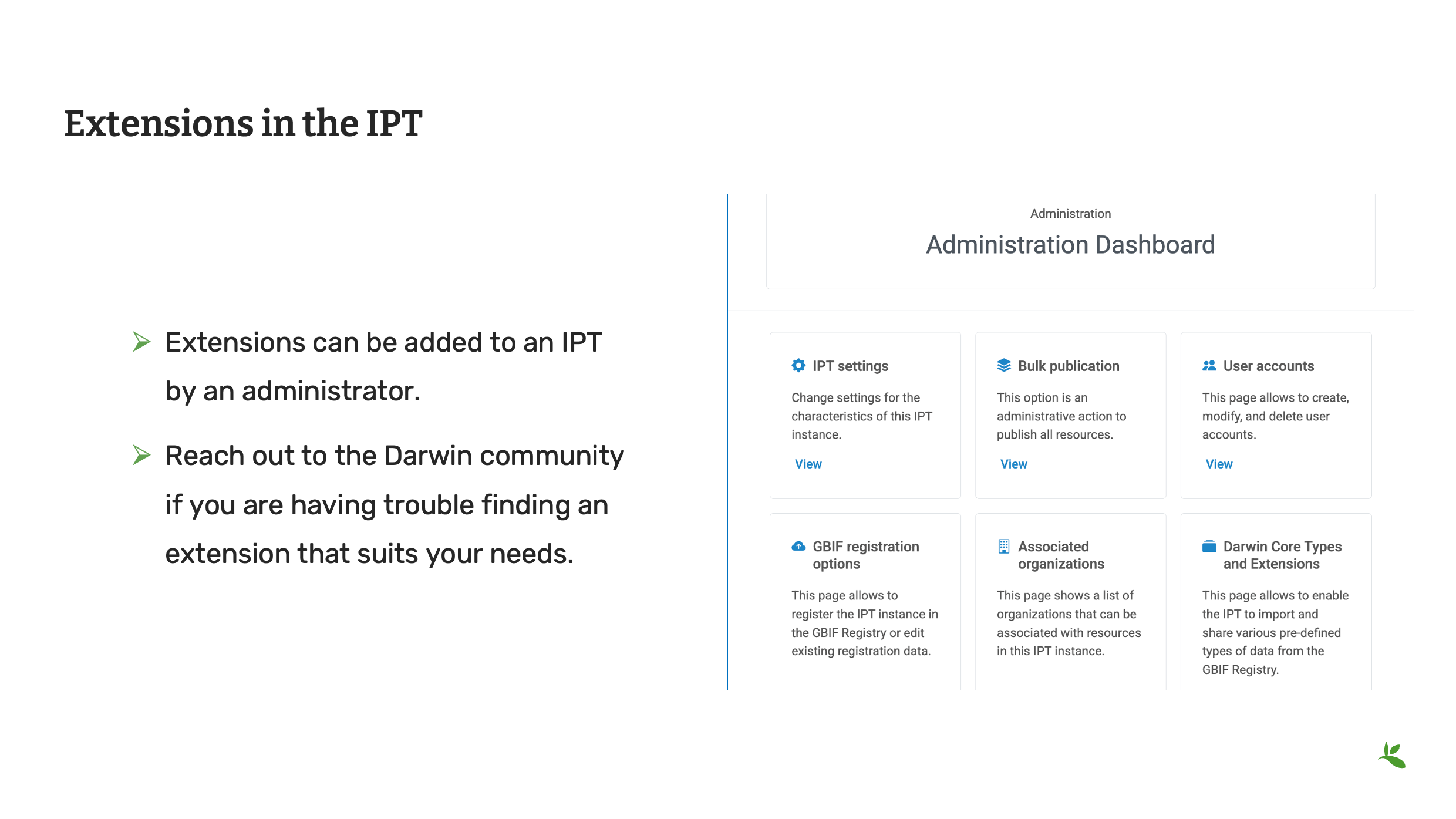

Slide 11 - Extensions in the IPT
Extensions were introduced in the Standards presentation. Publishing a dataset is now where you will put them into use. As mentioned previously, GBIF maintains the list of approved and draft extensions on its tools subsite. The approved extensions can be enabled by an IPT administrator on a Production IPT. Draft extensions can be enabled by an IPT administrator on a Test IPT. If you know you have data that cannot be shared with Simple Darwin Core, it is worth reviewing the list of available extensions.
If you think an extension doesn’t exist, try reaching out to members of the TDWG community or posting on the Darwin Core issues site. Community members will help guide you.
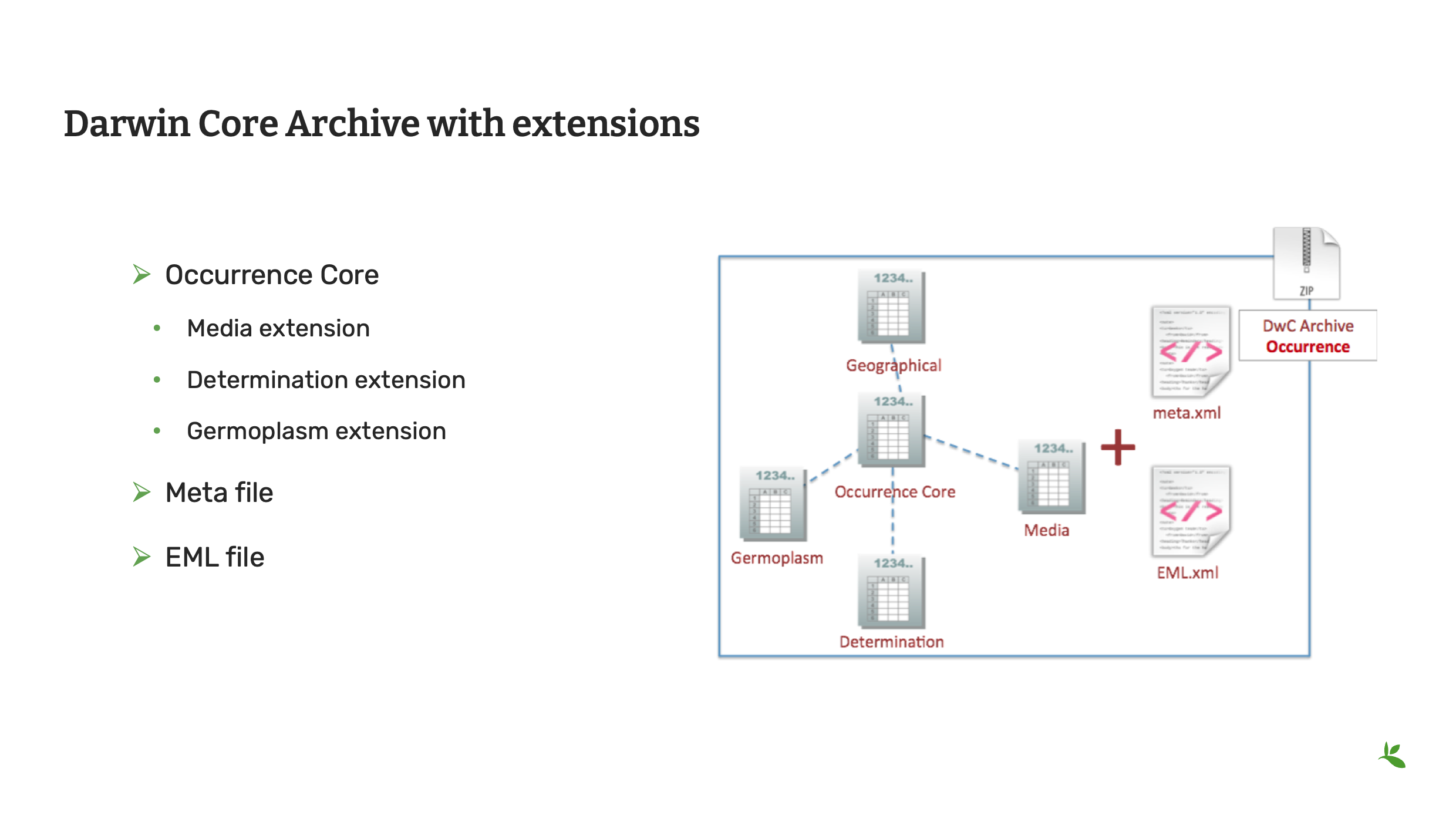
Slide 12 - Darwin Core Archive with extensions
Also, in the standards presentation, you were introduced to Darwin Core Archives. In the near future we will begin preparing the community to consider or otherwise adopt the Darwin Core Data Package, but for now we will continue with Darwin Core Archives.
This is an example of a Darwin Core archive with extensions.
It’s an occurrence dataset that has been published with images, determination history and genebank information.
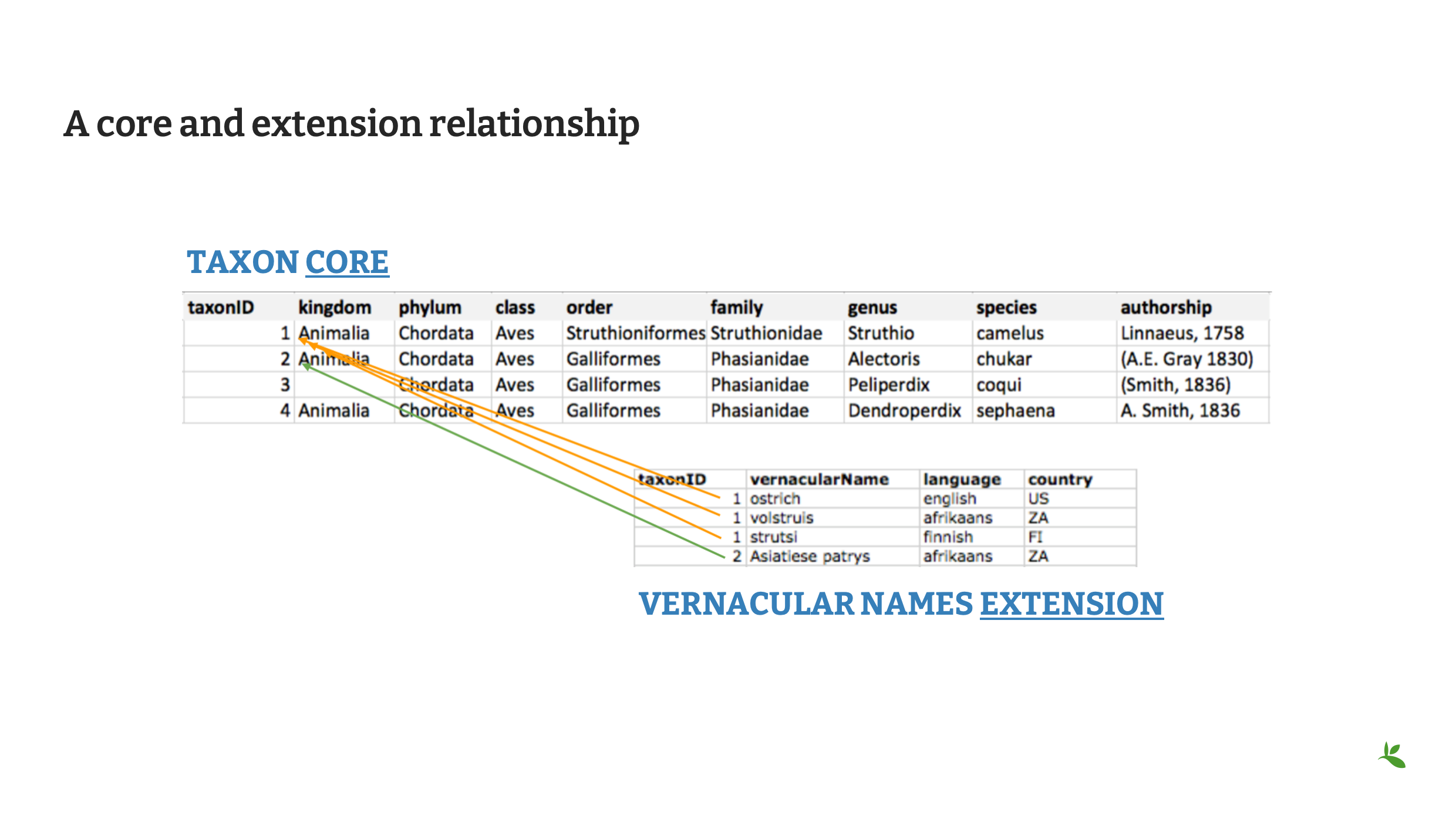
Slide 13 - A core and extension relationship
Until now, it may have been difficult to imagine how two single files can be related. A core with an extension can represent the following relationships.
-
1:0
-
1:1
-
1:many
The core file will always contain unique records. In this example using the Taxon Core, each taxon is unique and is represented by a taxonID. In Vernacular Names extension file, each vernacular name is unique. The first three represent common names for the species Struthio camelus (1:many relationship). The fourth name represents the common name for the species Alectoris chukar (1:1 relationship).
In this example the third and fourth taxons in the core file, do not have a match in the extension file. (1:0 relationship).
Slide 14 - Registering as a data publisher with GBIF
As we finish up this presentation, we’d like like to discuss registering as a data publisher at GBIF. The steps for registering were covered in the Introduction to GBIF course.
Data holding institutions are listed as the data publisher when their data is shared with GBIF. This allows users to properly provide credit and attribution for the data/datasets that they intend to use for research. And once users begin to site the dataset, the publishers page on GBIF.org will link back to literature citing the dataset.

Slide 15 - Aims of the publisher endorsement process
Before GBIF indexes a new publisher’s datasets, the institution must receive endorsement as a data publisher from one of the Participant nodes that coordinate activities of the national and organizational Participants in the GBIF network. If your country is not yet a participant, GBIF will coordinate the endorsement request through the GBIF Nodes Steering Group (NSG).
It is important to complete this step early in the publishing process as it could take up to a month for your publisher record to be endorsed by a Participant Node or the Nodes Steering Group.
La procédure d’endossement vise à garantir que :
-
Les données publiées sont pertinentes pour la portée et les objectifs du GBIF
-
Arrangements for data hosting are stable and persistent
-
La publication et l’utilisation des données sont soutenues par un engagement fort, national, régional et thématique
-
Data are as open as possible and available for sharing and reuse
-
Les éditeurs de données peuvent répondre aux commentaires et améliorer la qualité des données
Once a publisher receives endorsement, they can begin registering their published datasets with GBIF. You’ll hear more about this during the IPT demonstration.

Slide 16 - Conclusion
This presentation is part of a series of presentations used in the GBIF Biodiversity Data Mobilization course. The biodiversity data mobilization curriculum was originally developed as part of the Biodiversity Information Development Programme funded by the European Union. This presentation was originally created by Sophie Pamerlon with additional contributions by Nicolas Noe, Laura Anne Russell and Dag Endresen, BID and BIFA Trainers, Mentors and students. Narration is by me, Melissa Liu.
IPT overview and demonstration
|
During this demonstration, you will receive an overview of the IPT data publishing interface and you will learn how to publish an occurrence dataset using an IPT. |
A live demonstration will be provided. Visit the IPT user manual for full documentation.
Exercice 3
|
In this exercise, you will publish an event dataset using the IPT. |
Publication de données
After cleaning the data in the dataset, the team considers that publishing the data online through GBIF could be a good way to make this effort visible. They plan on assigning the data with a CC0 waiver. You have been requested to lead that publishing work.
-
Download UC-Practice-3-ForPublication.xlsx. (22 KB)
-
Open the file in Excel and export each tab as an individual CSV file (resulting in 4 files for upload to the IPT).
-
Use the assigned IPT installation to publish the dataset and register it with GBIF.
-
Utilisez la feuille d’exercice précédemment téléchargée pour donner vos réponses.
Révision
|
Testez vos connaissances sur les concepts abordés dans ce module. Certaines questions peuvent comporter plusieurs réponses correctes. Vous pouvez en savoir plus sur les réponses dans le Annexe des solutions. |