Capture de données
|
Dans ce module, vous découvrirez le concept de normes, et plus particulièrement la norme Darwin Core et ses composantes. Vous découvrirez également les différents types de données primaires sur la biodiversité et comment partager au mieux ces informations au sein du GBIF. Enfin, vous passerez en revue les principes de qualité des données dans le contexte de la capture des données et vous en apprendrez davantage sur la qualité et la cohérence des données (en particulier sur des sujets tels que le géoréférencement, les dates, les noms et la vérification croisée des taxons). |
Standards and Darwin Core
|
In this video (23:25), you will learn how you interact with standards every day. Then you will be introduced to Biodiversity Information Standards, including the Darwin Core Standard with which you will continue to use throughout this course. Subtitles are not yet available for this video. If you are unable to watch the embedded video, you can download it locally. (MP4 - 71.5 MB) If you prefer to read, you will find the transcript below the embedded video. |
Transcription de la présentation
Cliquez pour développer

Diapositive 1 - Normes et Darwin Core
Dans cette présentation, nous vous présenterons les normes de données relatives à la biodiversité. Nous nous concentrerons plus particulièrement sur la norme Darwin Core.

Diapositive 2 - Normes : Mettons-nous d’accord pour être d’accord
L’ingénieur et industriel W. Edwards Deming a dit :
« La standardisation ne signifie pas que nous portions tous des vêtements de la même couleur et du même tissage, que nous mangions des sandwichs standardisés ou que nous vivions dans des pièces standardisées avec un mobilier standardisé. Des maisons d’une variété infinie de styles sont construites avec quelques types de briques, du bois de dimensions standardisées, et des canalisations d’eau et de chauffage ainsi que des accessoires aux dimensions standardisées ».
Ce qu’il essayait de dire, c’est que la standardisation ne nous empêche pas d’être créatifs. Il nous montrait aussi que les normes sont déjà omniprésentes dans notre quotidien.
Au fil de cette formation, nous définirons le terme "norme" et examinerons comment nous interagissons quotidiennement avec les normes. Nous vous présenterons ensuite les normes d’information sur la biodiversité, notamment la norme Darwin Core que vous utiliserez tout au long de ce cours.

Diapositive 3 - Qu’est-ce qu’une norme ?
Qu’est-ce qu’une norme ?
Dans sa forme la plus simple, c’est :
"Une façon convenue de faire quelque chose."
Les normes sont une combinaison de règles, de conventions, de spécifications, d’exigences et de restrictions.

Diapositive 4 - Normes quotidiennes
L’objectif principal des normes est de créer un cadre de « compréhension mutuelle ». Elles doivent apporter de la clarté et faciliter la communication.
Voici quelques exemples de choses du quotidien qui utilisent des normes pour faciliter la communication d’informations :
-
Unités de mesure
-
Systèmes numériques
-
Alphabets
-
Langues
-
Émojis
-
Adresses postales
-
Code Morse
-
Codes-barres
Image de Mesure par Arek Socha depuis Pixabay https://pixabay.com/photos/measurement-millimeter-centimeter-1476913/ Image de Nombres par Clker-Free-Vector-Images depuis Pixabay https://pixabay.com/vectors/numerals-roman-number-blocks-35937/ Image d’Alphabet par https://en.wikipedia.org/wiki/Alphabet#/media/File:Historical_Writing_Systems_Template_Image.jpg Image de Langues par Christopher Kuszajewski depuis Pixabay https://pixabay.com/photos/source-code-code-programming-c-583537/ Image de Code Morse par https://en.wikipedia.org/wiki/Morse_code#/media/File:International_Morse_Code.svg Image de Courrier par OpenClipart-Vectors depuis Pixabay https://pixabay.com/vectors/airmail-mail-envelope-post-postal-145212/ Image d’Emojis par https://en.wikipedia.org/wiki/Emoticon#/media/File:Emoticon_Smile_Face.svg Image de Code-barre par IBOL https://ibol.org/site/wp-content/uploads/2018/12/bookmark-vectors.png

Diapositive 5 - Norme quotidienne - Un exemple
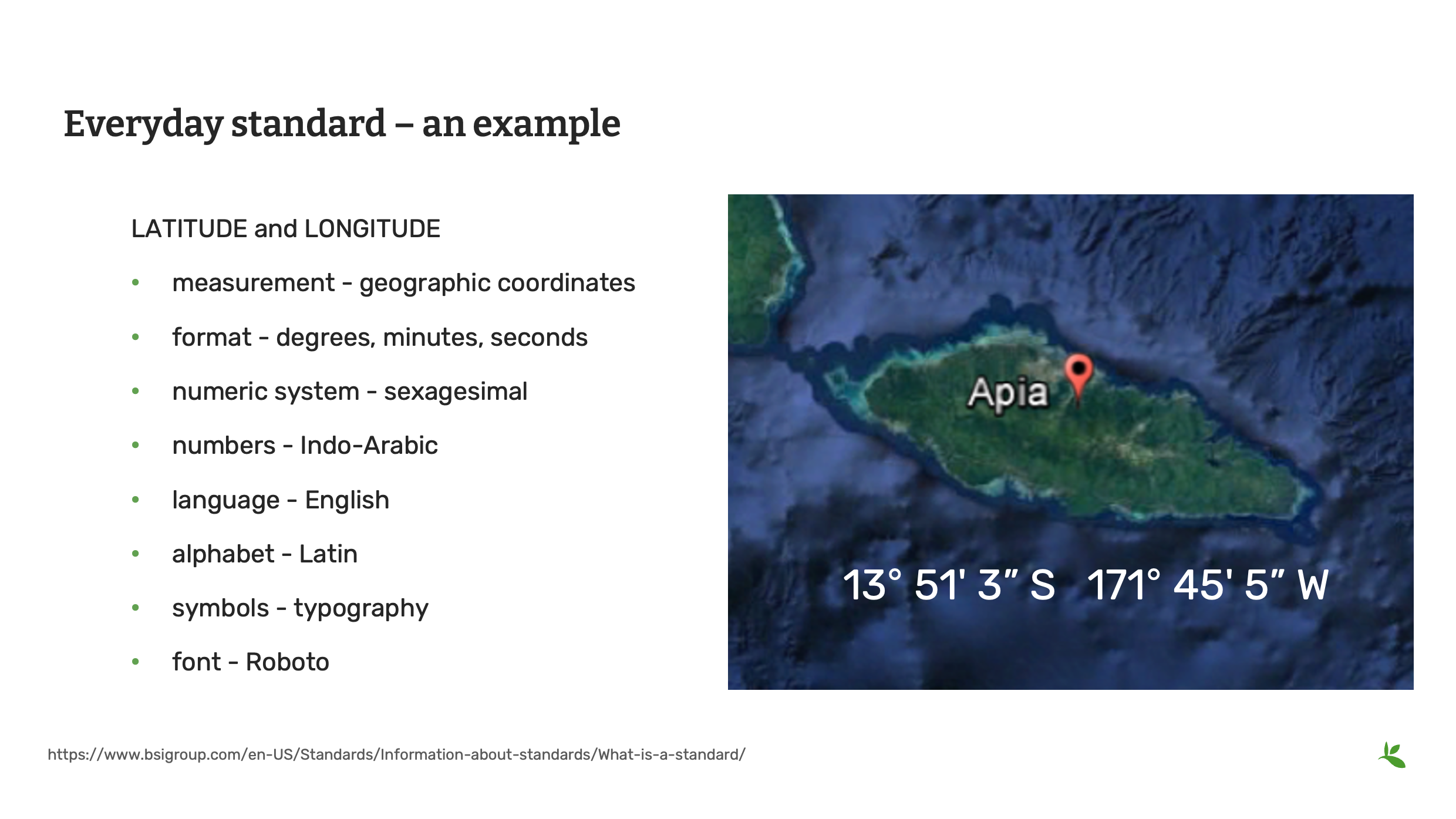
Prenons un exemple très précis et analysons-le. Ici, pour communiquer avec précision et de manière répétable une position sur Terre – une latitude et une longitude – il faut en fait combiner au moins 8 normes.
mesure - format des coordonnées géographiques - système numérique degrés, minutes, secondes - nombres sexagésimaux - langue indo-arabe - alphabet anglais - symboles latins - police de caractères typographiques - Roboto

Diapositive 6 - Règles et restrictions
Les normes permettent de restreindre l’éventail des possibilités. Dans les présentations précédentes sur les fondements du traitement de données, vous avez découvert les types de données, les schémas, les formats et les encodages de caractères. Chacun de ces éléments peut être utilisé pour restreindre l’éventail des possibilités dans le cadre d’une norme.
Les types de données peuvent restreindre les valeurs d’un champ. Ainsi, le texte alphanumérique va dans un champ de texte, les nombres décimaux dans un champ de nombre à virgule flottante. Vrai/Faux dans un booléen.
Un schéma d’encodage peut restreindre la plage de valeurs dans un champ. Par exemple, la liste de valeurs de latitude est possible dans une certaine plage : entre -90 et 90.
Un format peut restreindre la représentation d’une donnée dans un champ. Par exemple, une donnée peut apparaître comme année mois jour, jour mois année, ou mois jour année.
Finalement, le codage des caractères fournit les règles d’interprétation des octets de données. Pour nos besoins, nous utiliserons UTF-8.
Photo : Eurema blanda (Boisduval, 1836) Observé au Nepal par Bird Explorers (http://creativecommons.org/licenses/by-nc/4.0/)

Diapositive 7 - Normes pour le transfert de données
Au cours de cette formation, vous apprendrez à partager vos données. A ce titre, vous rencontrerez des normes de transfert de données.
Un schéma d’application permet de combiner des normes de données dans un but spécifique. Par exemple, utiliser les termes du Darwin Core dans les archives du Darwin Core. Nous allons approfondir ces deux types de schémas dans quelques instants.
Une fois de plus, vous utiliserez un format, mais cette fois le format limite les structures du jeu de données. Un jeu de données peut utiliser csv, xml, json et rdf.
Enfin, vous disposerez d’un protocole de transfert, qui fournit des informations sur la manière et l’endroit où envoyer des informations. Il peut s’agir de http (protocole de transfert hypertexte), ftp (protocole de transfert de fichier) et smtp (protocole simple de transfert de courrier ou, si vous préférez : envoyer du courrier à des personnes).
Image par Gerd Altmann depuis Pixabay https://pixabay.com/illustrations/web-network-computer-digital-5371553/

Diapositive 8 - Normes d’information sur la biodiversité
Dans le domaine de l’informatique de la biodiversité, il existe déjà de nombreuses normes qui peuvent vous aider à travailler avec vos données. L’USGS en donne une définition très précise :
"Les normes de données sont les règles selon lesquelles les données sont décrites et enregistrées. Afin de partager, d’échanger et de comprendre les données, nous devons standardiser leur format et leur signification.
Le résultat de l’utilisation de ces normes, lorsque approprié, permet d’augmenter l’intégrité, la précision et la cohérence des données en clarifiant les significations ambiguës et en minimisant les données redondantes.
Celles que vous êtes susceptible de rencontrer ou d’utiliser régulièrement peuvent inclure :
Ecological Metadata Language Standard (EML) Humboldt Ecological Inventory (Humboldt extension) Global Genome Biodiversity Network (GGBN) Ocean Data Standards and Best Practices Project (ODSBP)
Enfin, nous consacrerons le reste de cette discussion au Darwin Core. La norme Darwin Core vous permettra de partager vos jeux de données d’occurrences, taxonomiques et d’événements.
Références : EML - https://knb.ecoinformatics.org/#external//emlparser/docs/index.html Audubon - https://terms.tdwg.org/wiki/Audubon_Core GGBN - http://wiki.ggbn.org/ggbn/GGBN_Data_Standard ODSBP - https://www.iode.org/index.php?option=com_content&view=article&id=369&Itemid=100083 DwC - http://rs.tdwg.org/dwc/
Diapositive 9 - Qu’est-ce que le Darwin Core
Le Darwin Core est une norme de biodiversité développée par la communauté de l’Informatique de la Biodiversité. Il a été initialement développé sous le Taxonomic Databases Working Group ou T D W G, prononcé TDWG. Ces dernières années, le groupe a été renommé Biodiversity Information Standards. Mais l’acronyme persiste car la communauté est très attachée au nom TDWG.
"La norme comprend un glossaire de termes (dans d’autres contextes, ces termes pourraient être appelés propriétés, éléments, champs, colonnes, attributs ou concepts) destinés à faciliter le partage d’informations sur la diversité biologique en fournissant des identificateurs, des étiquettes et des définitions. Le Darwin Core est principalement basé sur les taxons, leur présence dans la nature telle que documentée par des observations, des spécimens, des échantillons et des informations connexes".
En résumé, le Darwin Core est une
"Liste des champs et de leurs définitions, relatifs aux données sur la biodiversité."
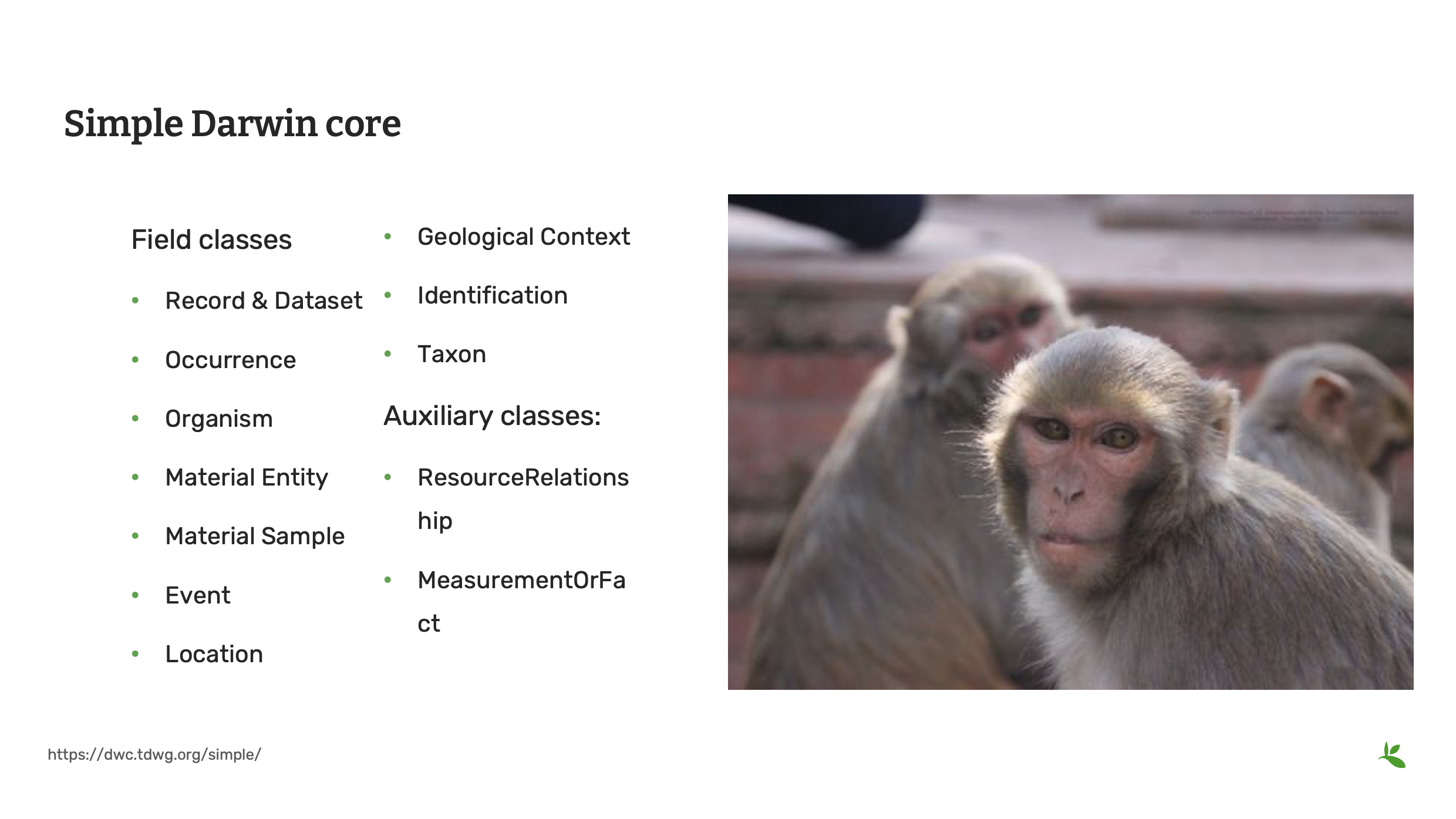

Diapositive 10 - Simple Darwin Core
Au fur et à mesure que nous approfondissons le Darwin Core ou, en abrégé, le DwC, vous apprendrez qu’il s’agit de bien plus que JUSTE une liste de champs. Nous utiliserons le Simple Darwin Core, qui est un sous-ensemble prédéfini de termes d’usage courant dans une grande variété d’applications de biodiversité.
Ce sous-ensemble contient plus de 150 champs, qui sont placés dans un ensemble de classes comprenant :
Enregistrement et Jeux de Données d’Occurrence Organisme Matériel d’Entité Matériel d’Échantillon Événement Lieu Contexte géologique Identification Taxon
De plus, il existe deux classes auxiliaires appelées :
ResourceRelationship MeasurementOrFact
D’après le guide de l’utilisateur du Simple Darwin Core, "le Simple Darwin Core est simple en ce sens qu’il ne suppose (et n’autorise) aucune structure au-delà du concept de lignes et de colonnes, que l’on peut considérer comme des attributs et leurs valeurs, ou des champs et des enregistrements".
Photo : Macaca mulatta (Zimmermann, 1780) Observé au Népal par Vladimir Tkalčić (http://creativecommons.org/licenses/by-nc/4.0/)
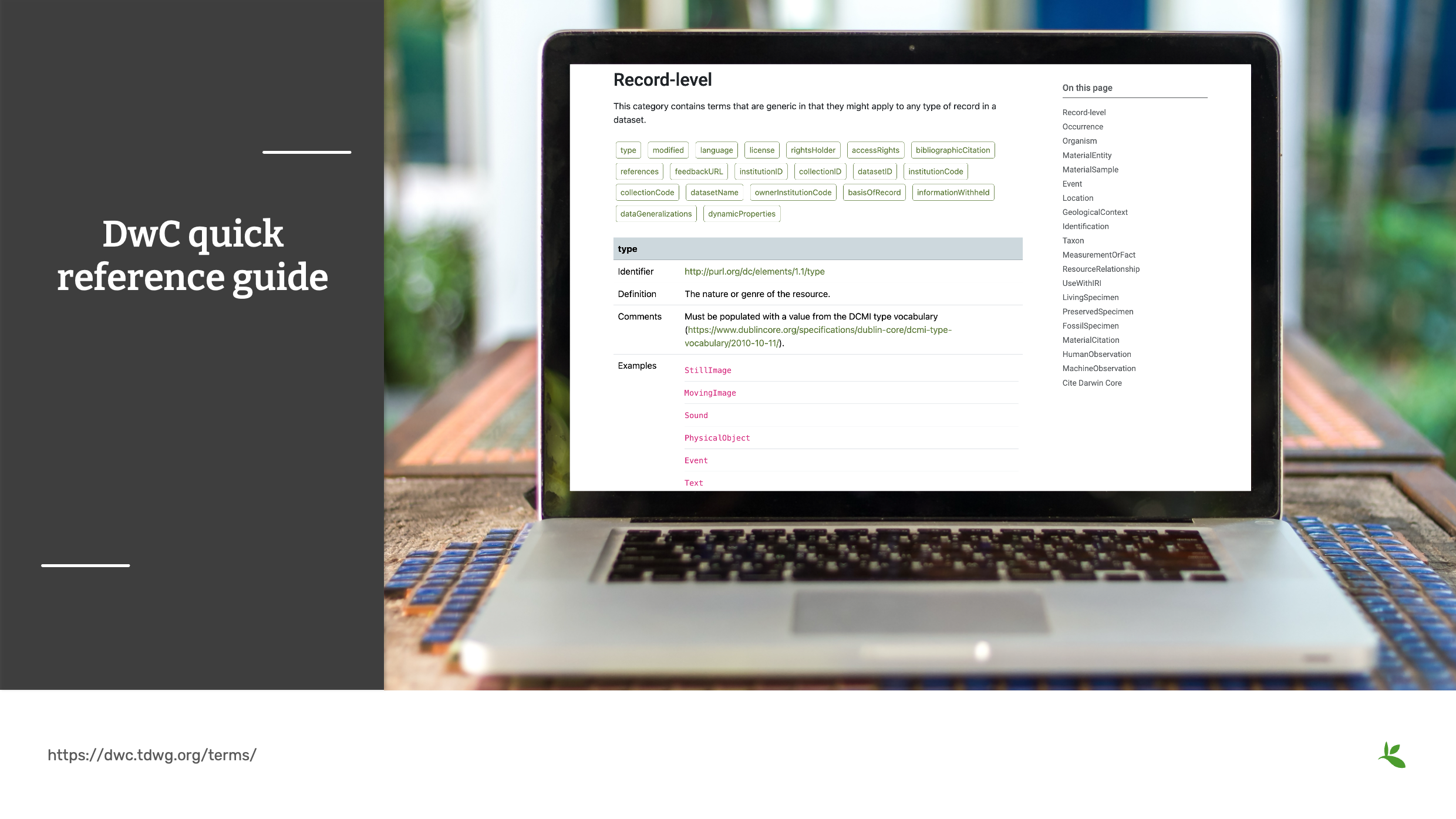
Diapositive 11 - Guide de référence rapide du Darwin Core
Le guide de référence rapide du DwC deviendra bientôt votre ressource préférée. Cette page fournit une liste de tous les termes actuellement recommandés dans la norme Darwin Core. Les catégories telles que Occurrence ou Event correspondent aux classes Darwin Core qui regroupent d’autres termes.

Diapositive 12 - Termes du DwC relatifs au pays et au code de pays
Nous allons maintenant examiner quelques exemples de termes du Darwin Core. Le guide de référence rapide présente chaque terme de manière cohérente avec le nom de l’identifiant, la définition, les commentaires et les exemples. Les premiers termes que nous examinerons sont Country et CountryCode dans la catégorie Location.
En général, les détenteurs de données disposent d’un champ pour le pays dans leurs données sources. Mais souvent, ces données peuvent être assez confuses, avec des fautes d’orthographe, des abréviations et des noms historiques. Il s’agit toutefois de l’un des éléments de données les plus faciles à standardiser. Comme indiqué dans les commentaires, la meilleure pratique recommandée est d’utiliser un vocabulaire contrôlé tel que le Getty Thesaurus of Geographic Names. Un vocabulaire contrôlé impose des restrictions sur les valeurs à utiliser pour ce terme.
CountryCode est un terme qui n’est généralement pas présent dans les données sources. Mais il s’agit là encore d’un champ qui peut facilement être alimenté en données grâce à la recommandation formulée dans les commentaires d’utiliser un code de pays ISO 3166-1-alpha-2.
Le GBIF recommande fortement le partage du champ CountryCode dans les jeux de données d’occurrence. Le partage du champ Country est également encouragé.
Vous en apprendrez davantage sur les exigences et les recommandations du GBIF dans les prochaines sessions.
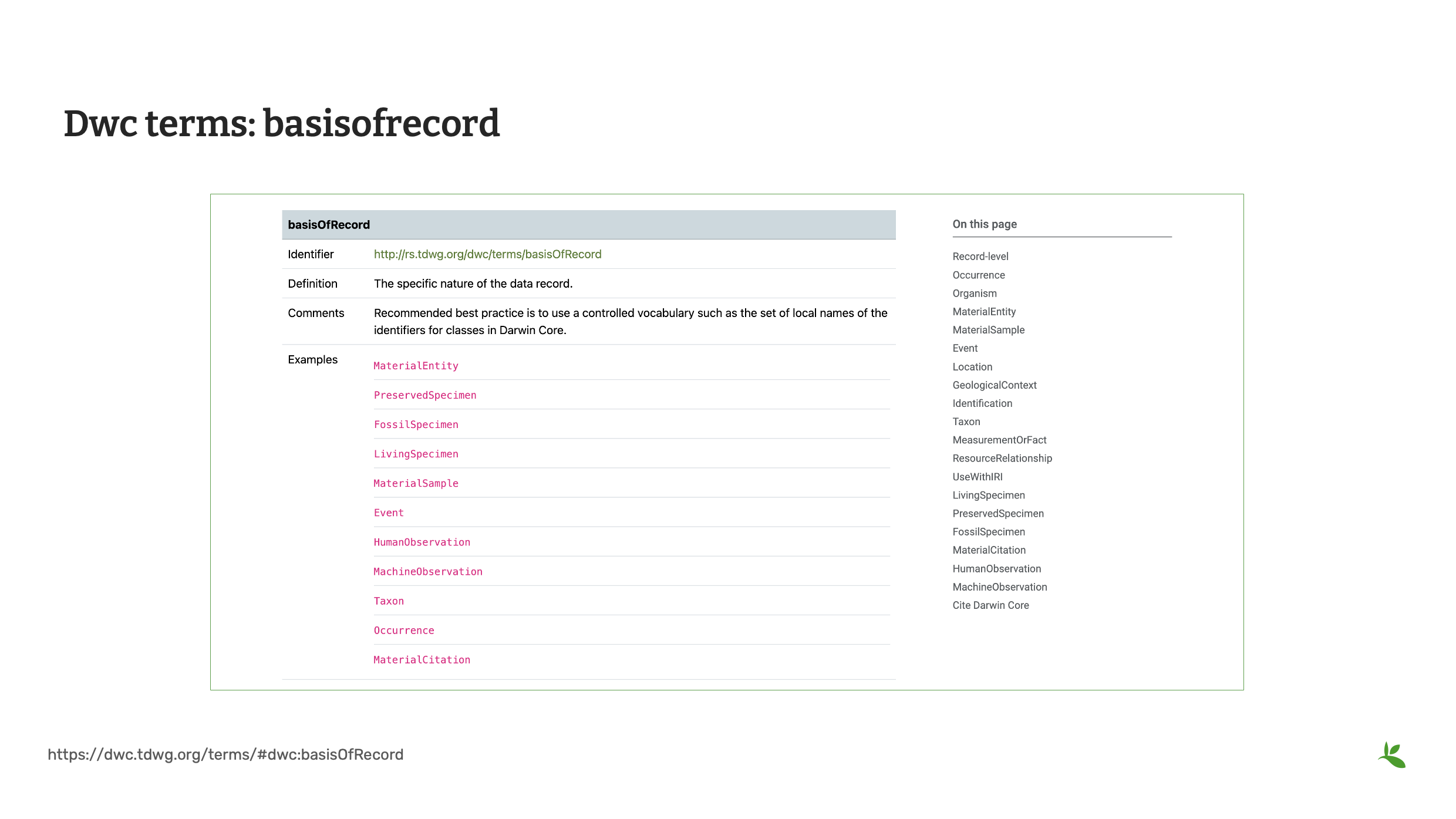
Diapositive 13 - Termes du DwC basisOfRecord
Le terme suivant est basisOfRecord. basisOfRecord définit la nature de chaque enregistrement dans un jeu de données. BasisOfRecord suit un vocabulaire contrôlé. Vous pouvez choisir entre PreservedSpecimen, FossilSpecimen, LivingSpecimen, MaterialSample, Event, HumanObservation, MachineObservation, Taxon ou Occurrence. Le GBIF exige basisOfRecord pour les jeux de données d’occurrences publiés.
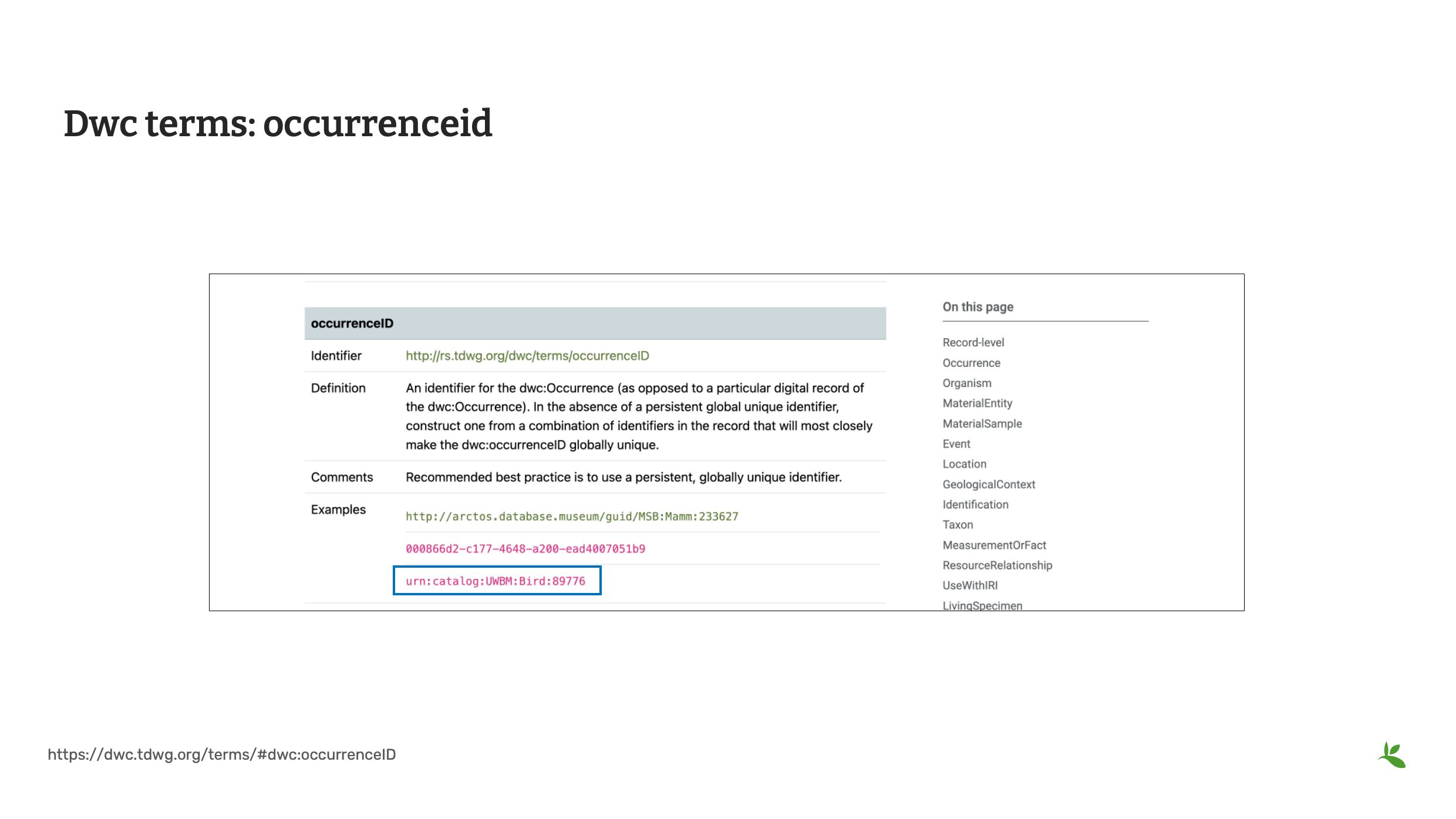
Diapositive 14 - Termes du DwC occurrenceID
Le dernier terme que nous allons examiner est occurrenceID. Lors de la publication d’enregistrements d’occurrence, le GBIF exige un identifiant d’occurrence (occurrenceID). Un occurrenceID est un identifiant pour l’occurrence elle-même, et non pour l’enregistrement numérique de l’occurrence. La meilleure pratique recommandée est d’utiliser un identifiant global unique, également connu sous le nom de GUID. En l’absence de GUID, un identifiant unique peut être composé d’autres identifiants dans le jeu de données. Il existe des outils sur Internet qui peuvent vous aider à générer des GUIDs pour vos enregistrements. Si vous utilisez cette méthode, ces GUIDs devraient devenir un champ permanent dans vos données sources identifiant chaque enregistrement. Pour l’exercice réalisé dans ce cours, vous créerez un occurrenceID avec un format similaire au troisième exemple dans la boîte bleue.

Diapositive 15 - Extensions du Darwin Core
En utilisant le Simple Darwin Core, vous découvrirez peut-être que vous avez d’autres données à partager mais que vous ne pouvez pas trouver les termes correspondants dans le DwC. Ces données peuvent être des images ou des fichiers sonores, ou peut-être êtes-vous responsable d’une collection de vertébrés et avez-vous compilé de nombreuses données sur le poids et la taille des spécimens. Ou même des informations historiques détaillées sur l’identification d’un taxon. Dans ce cas, vous vous tournerez vers les extensions du Darwin Core afin d’étendre les données de base en fournissant des fichiers supplémentaires qui correspondent aux données de base. Les extensions qui répondraient à vos besoins dans ces trois exemples sont les suivantes :
Simple Multimedia Measurements or Facts Identification History
Il existe de nombreuses autres extensions. Le GBIF tient à jour une liste de toutes les extensions approuvées et en projet sur son sous-site consacré aux outils.
Photo : Aleuria aurantia (Pers.) Fuckel Observé au Népal par Elizabeth Byers http://creativecommons.org/licenses/by-nc/4.0/
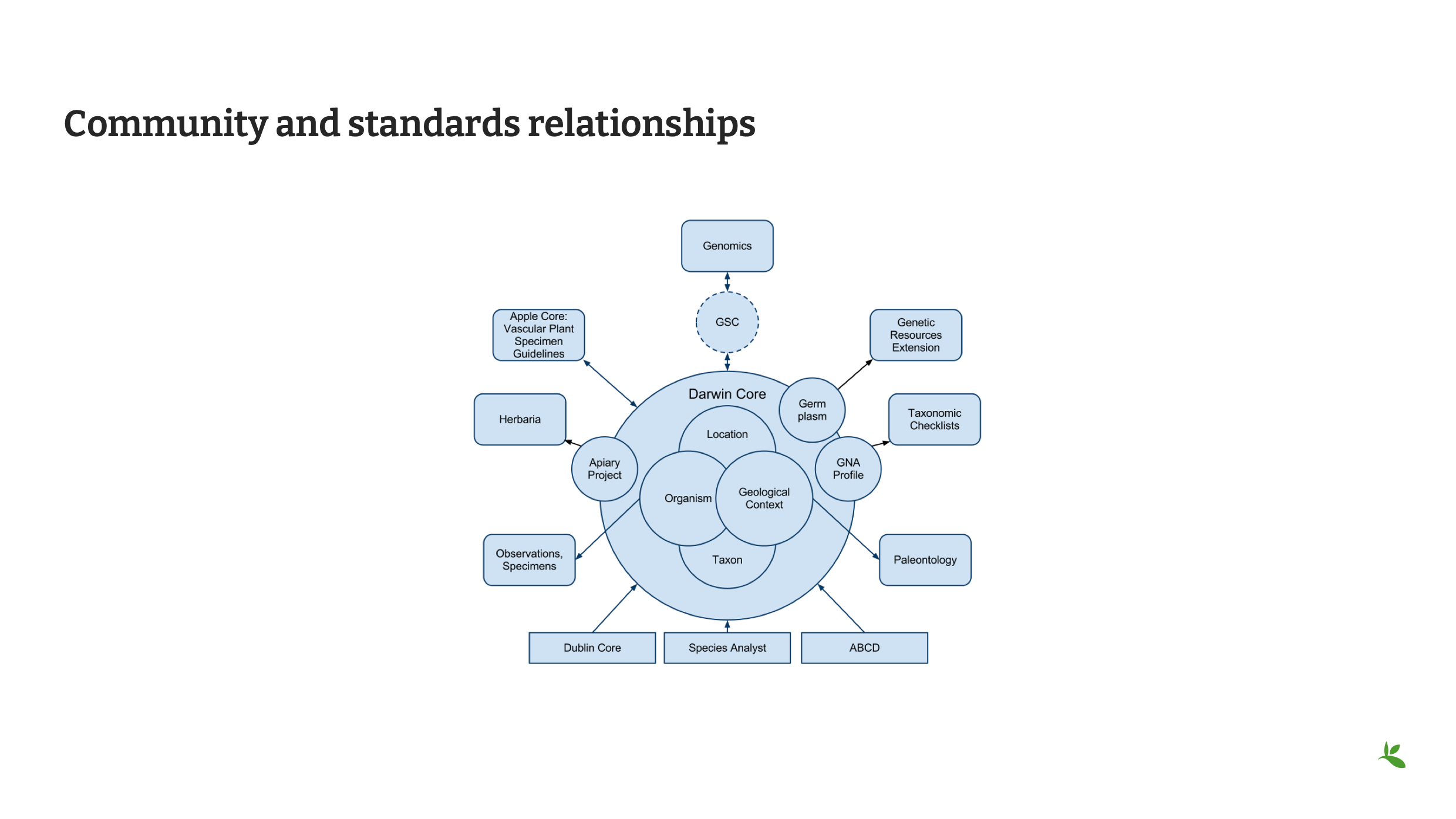
Diapositive 16 - Relations entre la communauté et les normes
Il existe de nombreuses couches dans notre communauté de l’informatique de la biodiversité. L’image montre ici les relations entre ces couches et où elles se recoupent avec le Darwin Core, ainsi que les extensions qui pourraient s’avérer nécessaires pour un partage complet des données.
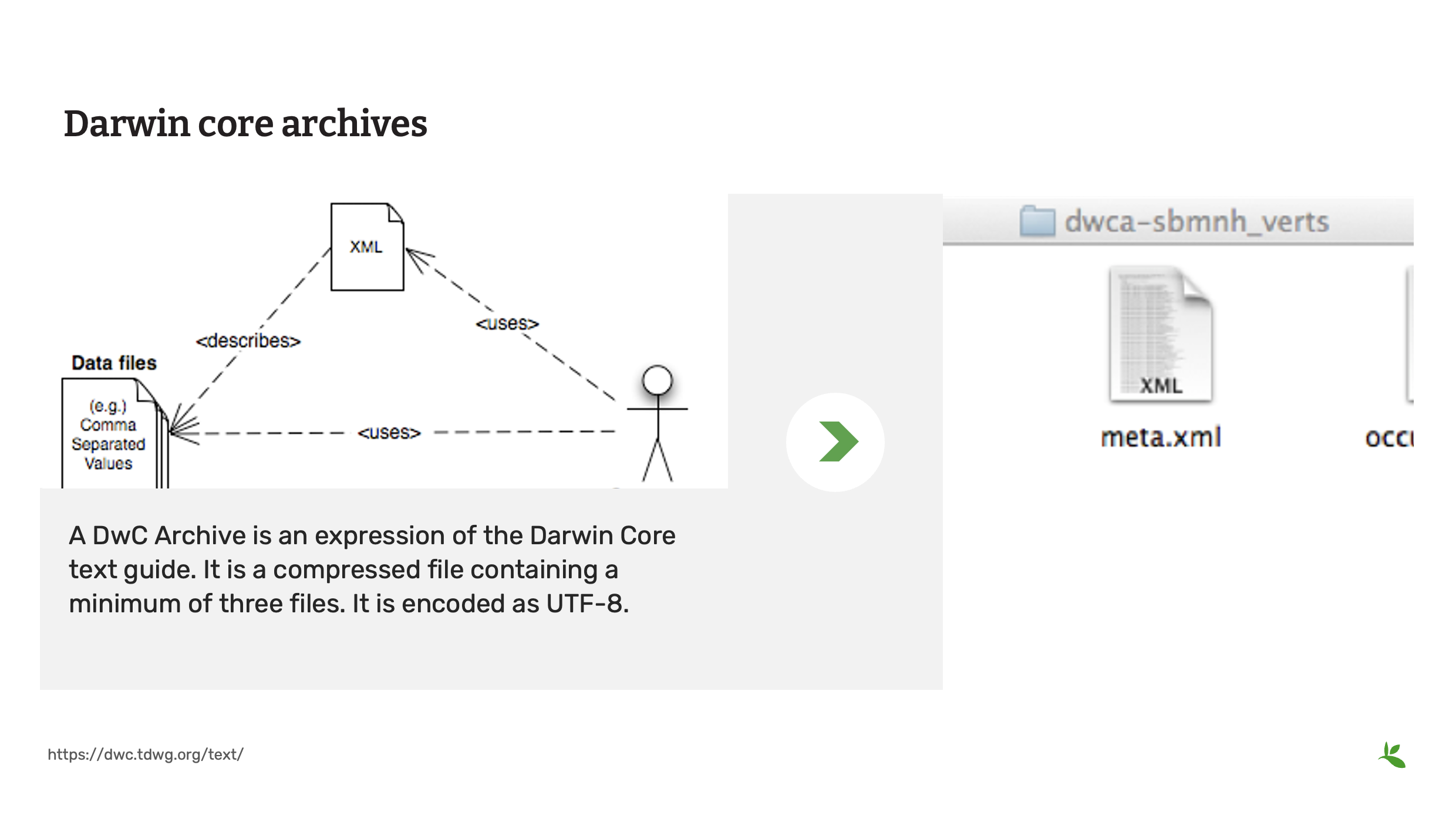
Diapositive 17 - Archive Darwin Core (DwC-A)
Les données partagées avec le GBIF sont actuellement soumises via une archive Darwin Core ou DwCA.
Un DwCA est une expression du guide textuel Darwin Core. Il s’agit d’un fichier compressé contenant au minimum trois fichiers. Il est encodé en UTF-8.
Dans cet exemple, ces trois fichiers sont :
Un fichier de données (occurrence.txt) conforme à la norme SIMPLEDWC au format CSV, dont la première ligne contient les noms des termes standard Darwin Core. Un fichier de métadonnées (meta.xml) au format XML contenant les informations techniques nécessaires à un ordinateur pour utiliser le fichier de données. Un fichier de métadonnées (eml.xml) au format XML contenant des informations explicatives sur les enregistrements contenus dans le fichier de données, afin d’indiquer à l’utilisateur si les données sont adaptées à son usage.
On peut obtenir une structure plus complexe en partageant plusieurs fichiers CSV liés afin d’enrichir les données. Ces fichiers sont liés au fichier principal par un identifiant unique. Dans un jeu de données d’occurrences, ces fichiers CSV sont liés par l’identifiant d’occurrence (occurrenceID).

Diapositive 18 - Mises à jour du DwC et passage au Darwin Core Data Package (DwC-DP)
Bien que les archives Darwin Core aient été notre méthode de publication privilégiée depuis 2012, nous nous orientons vers un nouveau modèle appelé Darwin Core Data Package.
Le nouveau modèle vise à "élargir l’éventail des questions scientifiques que le GBIF peut aborder."
It will allow us to expand our data scope, engage with new data communities, and build tools to enable data flows based on the updated standard.
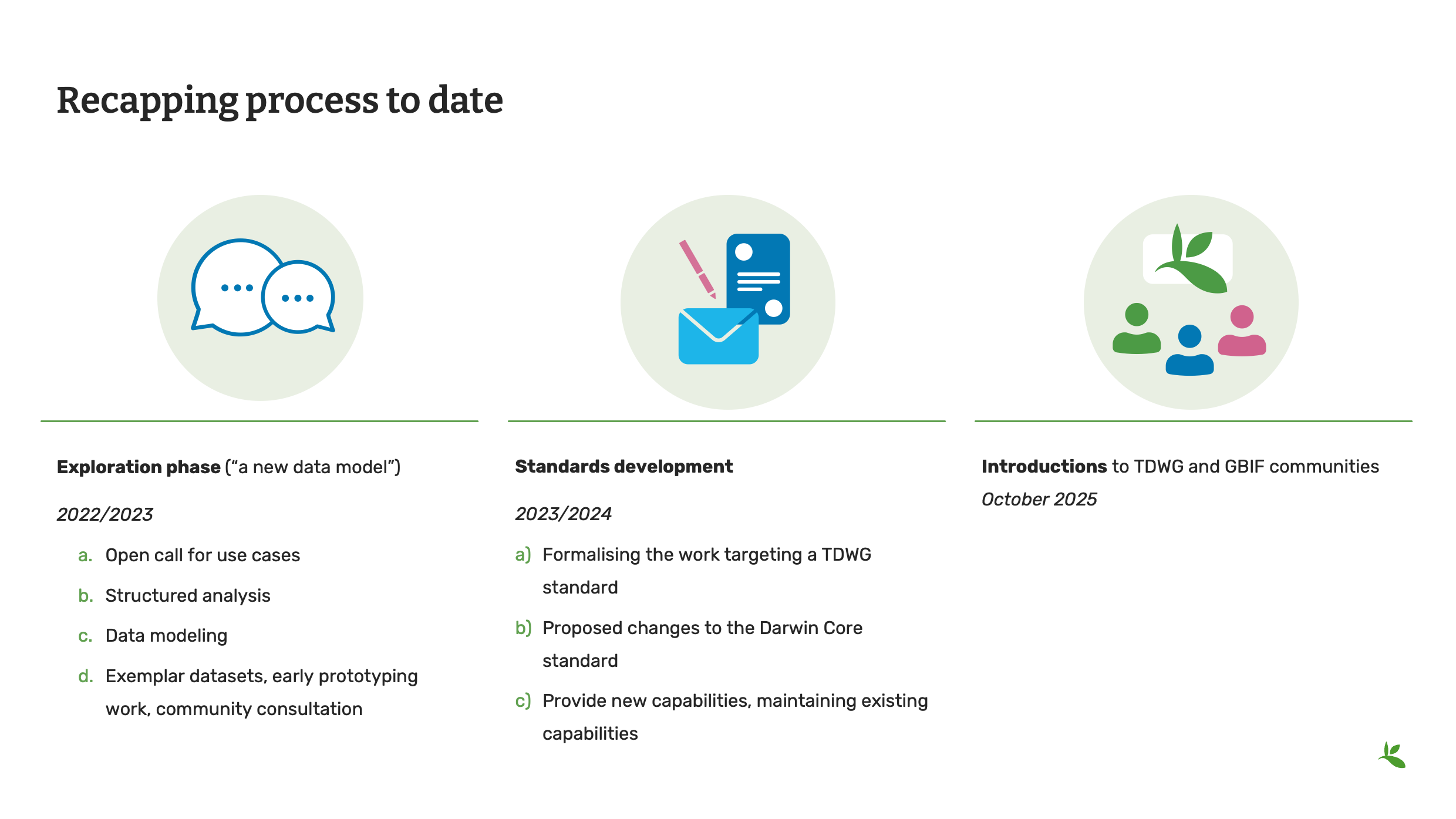
Diapositive 19 - Récapitulatif du processus à ce jour
La communauté travaille sur ce modèle depuis 2022 et les normes relatives à ce modèle sont en cours d’élaboration depuis 2023.
We are finally in the community review and ratification phase for the new terms and the new data package which was released in October of 2025. As you can see from this timeline, it takes a long time implement change to an established standard and it takes a dedicated community to see it through.

Diapositive 20 - Modifications proposées du Darwin Core
Les changements soumis à consultation publique comprennent :
65 nouveaux termes, 75 modifications proposées aux termes existants, un modèle conceptuel documenté pour les lignes directrices DwC relatives à l’utilisation du format de données sans friction (Frictionless Data Format)
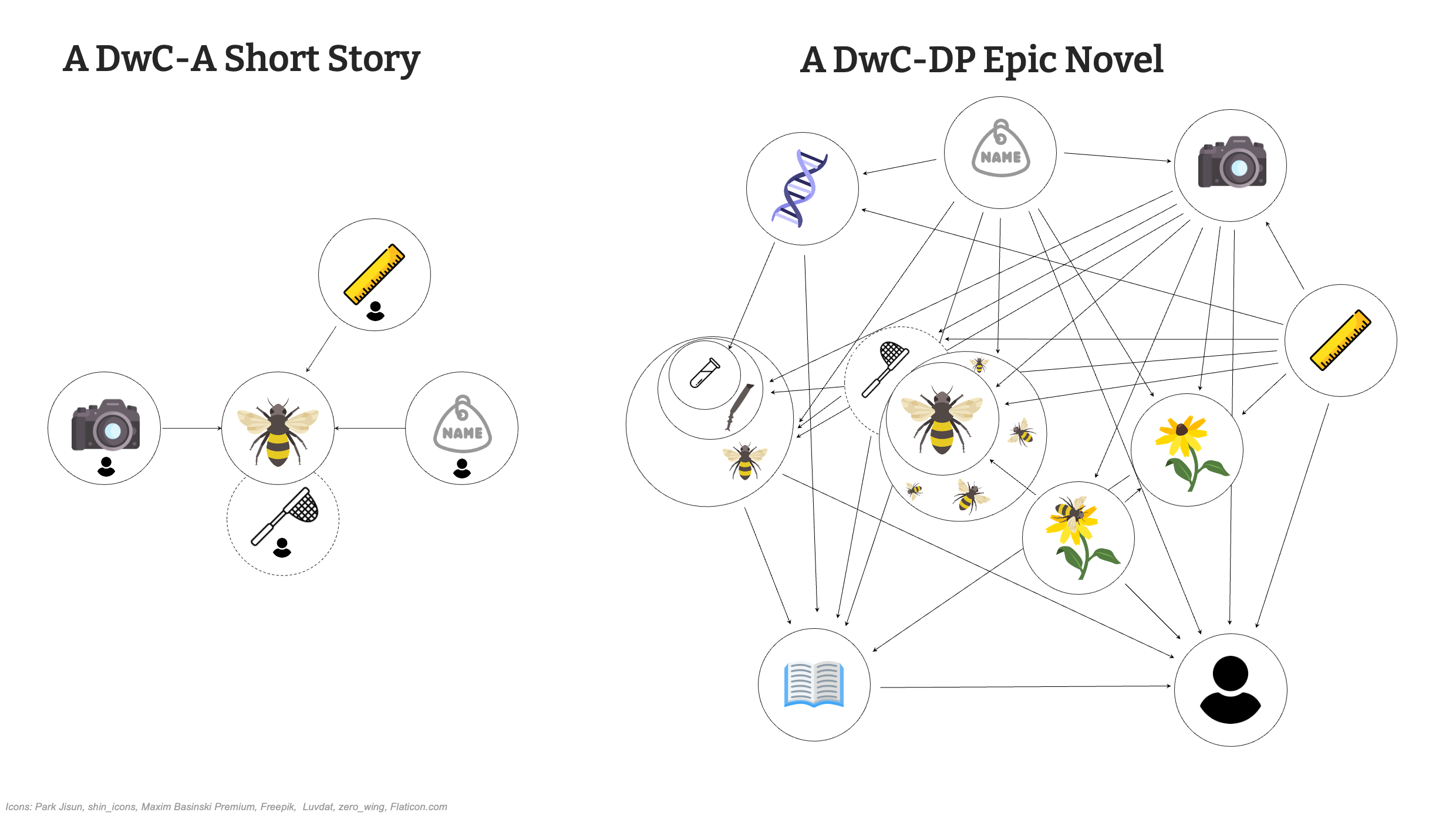
Diapositive 21 - Une nouvelle DwC-A OU un roman épique DwC-DP ?
Le Darwin Core Data Package ouvre complètement les possibilités de partage de données, transformant une nouvelle des archives Darwin Core en un roman épique basé sur le Darwin Core Data Package.
À titre de comparaison, on peut raconter des histoires avec DwC-A ou DwC-DP, mais il est évident que l’éventail des efforts scientifiques qui peuvent être fidèlement retranscrites par DwC-DP est beaucoup plus vaste.
Abeille, fleur : Park Jisun ADN : Luvdat Livre : zero_wing Filet : shin_icons Personne : Maxim Basinski Tube à essai premium : Freepik Règle : Freepik Appareil photo : Freepik Identification : Freepik

Diapositive 22 - Dois-je vraiment faire tout ça ?
Tout cela peut paraître déroutant lorsque l’on découvre la mobilisation des données, mais nous tenons à vous informer que des changements sont à venir qui, selon la nature de vos données, pourraient s’avérer très intéressants pour vous.
Nous reconnaissons également que tout le monde n’a pas besoin de faire tout cela, surtout au sein d’un seul ensemble de données, et que les éditeurs pourront n’utiliser que ce dont ils ont besoin.
Pour le moment, concernant ce cours, nous nous concentrons sur la formation utilisant le noyau événement (event core), à propos duquel vous découvrirez plus de détails lors des prochaines sessions.
Une fois le processus de ratification terminé et lorsque nous serons prêts à ce que les éditeurs commencent à utiliser le Darwin Core Data Package, nous organiserons des événements communautaires virtuels pour compléter la formation.
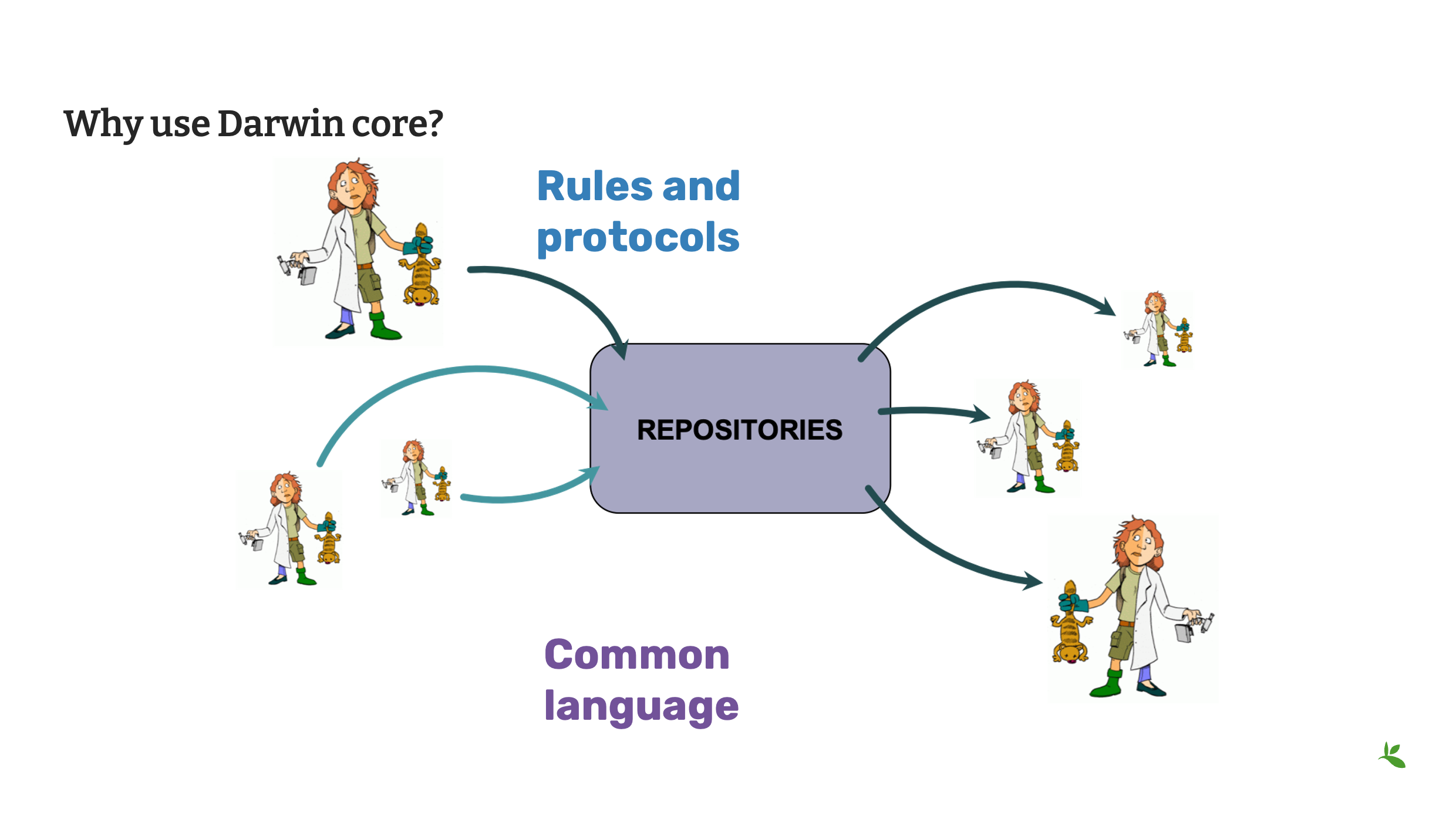
Diapositive 23 - Pourquoi utiliser le Darwin Core ?
En conclusion, nous avons couvert ce qu’est le Darwin Core et nous espérons que vous avez commencé à comprendre pourquoi vous devriez l’utiliser.
C’est une norme, et les normes sont une bonne chose ! Elles nous fournissent les règles et les protocoles nécessaires pour partager nos données avec les autres.
Le Darwin Core nous fournit également une langue commune. Comme nous l’avons vu dans la partie Fondations – Présentation de la documentation, les données sources peuvent être difficiles à exploiter lorsqu’on essaie de comparer les jeux de données. Les champs dans vos données sources peuvent être différents des champs dans des données source provenant d’une autre institution. Lorsque nous utilisons tous le Darwin Core pour partager nos données, nous comprenons que les données ont été partagées avec une langue commune.
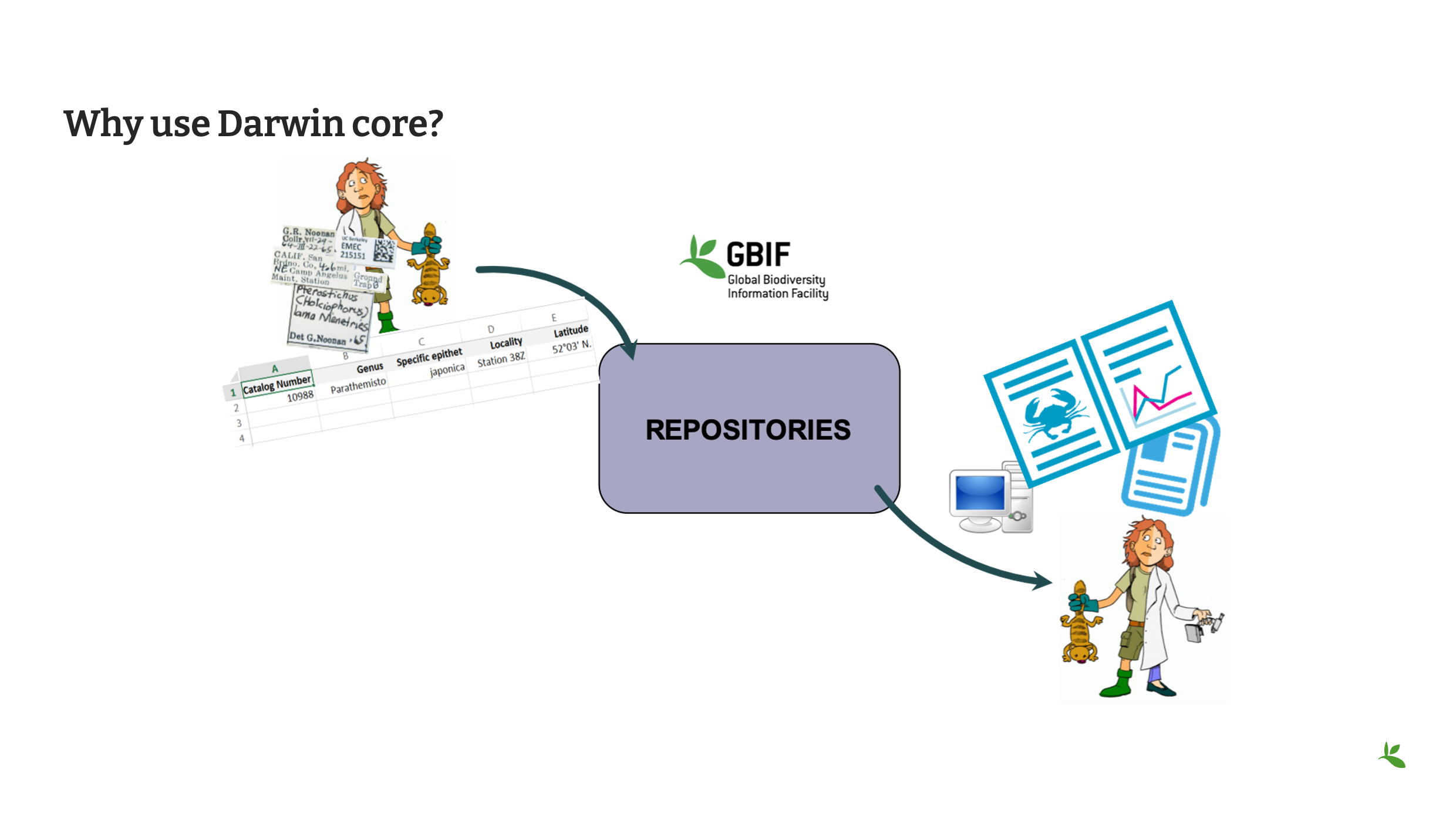
Diapositive 24 - Pourquoi utiliser le Darwin Core ?
Et ce ne sont pas seulement les détenteurs de données qui comprennent ce langage commun, mais aussi les utilisateurs de données. Après tout, qu’y a-t-il de mieux qu’un utilisateur qui trouve un jeu de données adapté à son usage, partagé dans un langage commun, qui lui permet de faire de progresser la recherche scientifique ?

Diapositive 25 - Conclusion
Ce document fait partie d’une série de présentations utilisées dans le cadre du cours sur la mobilisation des données sur la biodiversité du GBIF. Le programme de mobilisation des données sur la biodiversité a été développé à l’origine dans le cadre du programme de développement de l’information sur la biodiversité financé par l’Union européenne. Cette présentation a été créée par Paula Zermoglio et John Wieczorek avec des contributions supplémentaires de Sharon Grant, Sophie Pamerlon, Laura Anne Russell, Cecilie Svenningsen et Dag Endresen. Cette présentation a été narrée par Vijay Barve. .
Exercice 1a
|
Pour cette activité, vous examinerez les noms de champs textuels et les associerez aux termes du Darwin Core |
-
Trouvez le terme du Darwin Core sur https://dwc.tdwg.org/terms/ qui correspond le mieux aux noms des champs.
-
Téléchargez UC-Practice-exercise-sheet_EN.docx pour fournir vos réponses.
Types de jeux de données GBIF pour les données primaires sur la biodiversité
|
In this video (08:16), you will review primary biodiversity data that can be shared within GBIF. Subtitles are not yet available for this video. If you are unable to watch the embedded video, you can download it locally. (MP4 - 28.8 MB) If you prefer to read, you will find the transcript below the embedded video. |
Transcription de la présentation
Cliquez pour développer

Diapositive 1 - Types de jeux de données GBIF pour les données primaires sur la biodiversité
In this presentation, we will have a look at the different types of data that can be called ‘primary biodiversity data’ and shared within GBIF. These data can be complex and have different origins. We’ll see how they can be structured into one of the dataset classes accepted by GBIF.
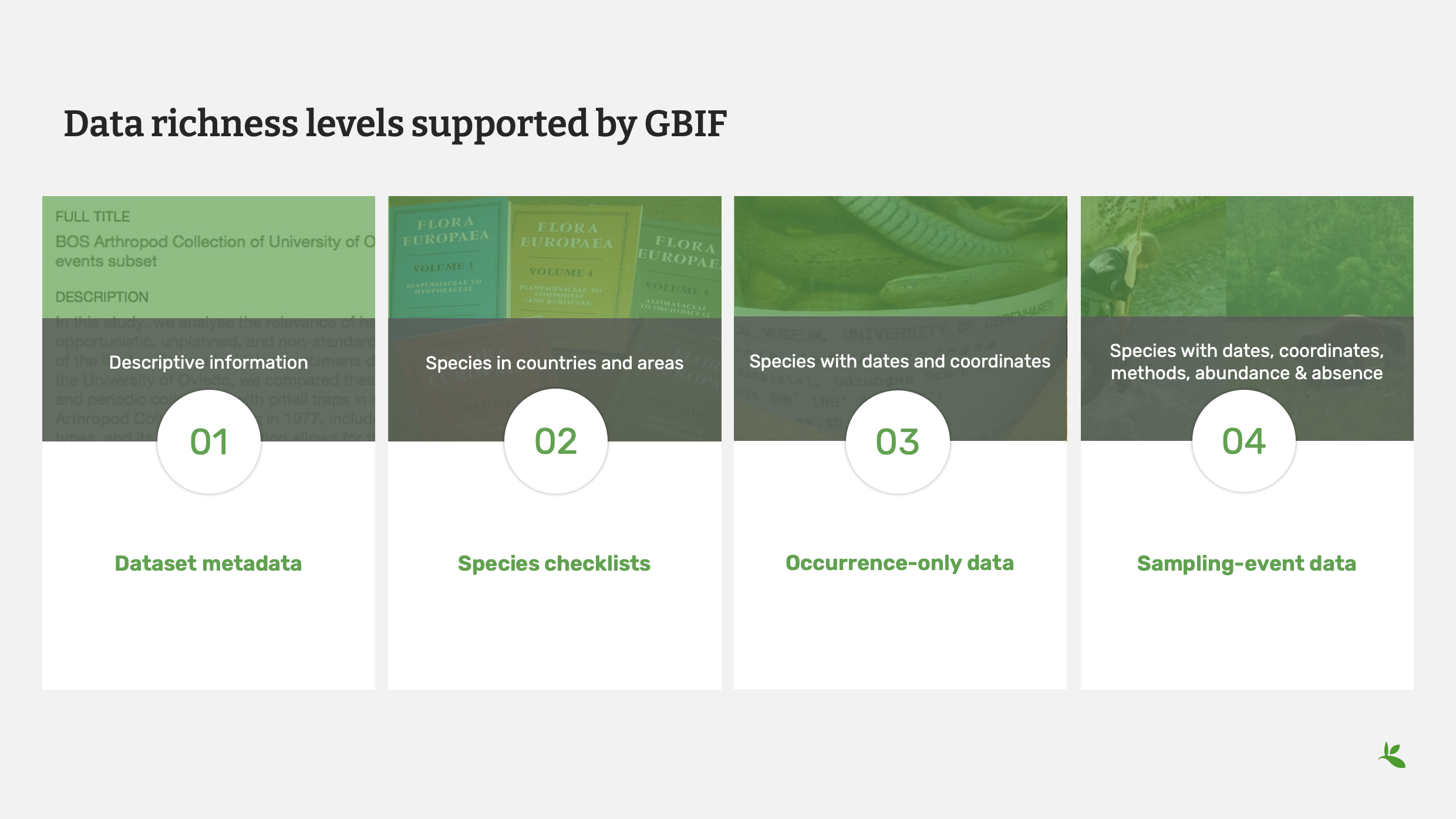
Diapositive 2 - Niveaux de richesse des données pris en charge par le GBIF
GBIF currently supports four types of datasets:
The first is dataset metadata.
This is a dataset that allows you to provide descriptive information about a dataset. You may use this type when you have not yet digitized your data. No data files are published with this type of dataset only descriptive metadata.
The second type are species checklists.
This allows you to share information on species including the countries and areas where they are found.
The third type is occurrence-only data.
This dataset type is for data that include names, dates and coordinates – the what, when and where of your data.
The last type is Sampling-event data.
This type allows you to share even more data. You can share species with dates, coordinates, methods, abundance and even absence.
We will now further explore the checklist, occurrence and sampling-event dataset types.

Slide 3 - Checklist data
Checklist datasets provide a catalogue or list of named organisms, or taxa. While they may include additional details like local species names or specimen citations, these ‘checklists’ typically categorize information along taxonomic, geographic, and thematic lines, or some combination of the three. For example, a dataset that catalogues the Red Listed molluscs of Seychelles has distinct elements of taxonomy (the phylum Mollusca), geography (the island nation of Seychelles) and theme (species deemed imperiled by IUCN experts). Checklists function as a rapid summary or baseline inventory of taxa in a given context.

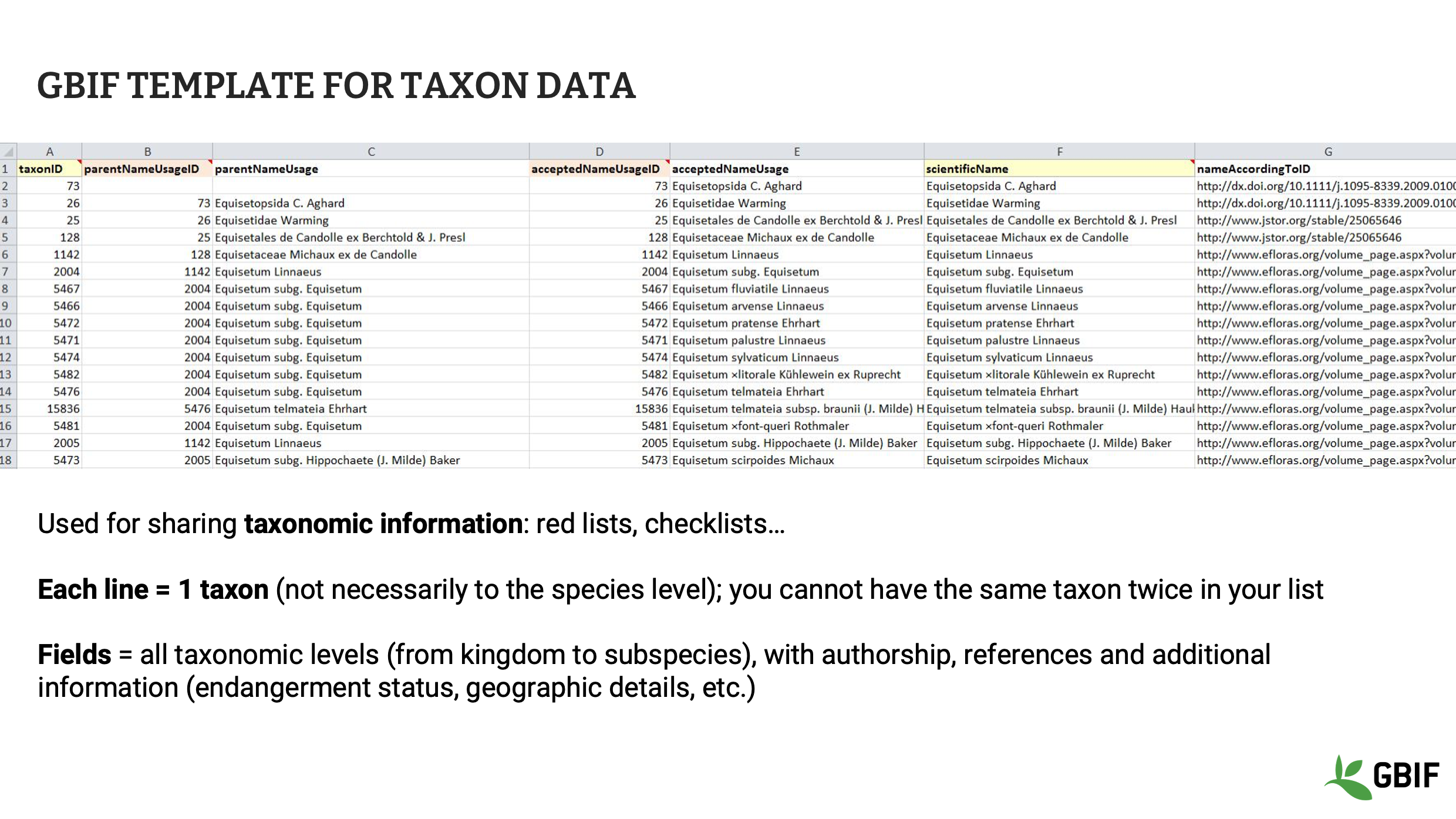
Slide 4 - GBIF template for taxon data
GBIF provides data publishers with templates for each class of dataset. The GBIF template for taxon data allows you to share information linked to each species or taxa including the taxon id, full name and authorship, relationship to other taxa, geographical details and so on. In this template, one line represents one taxon and a taxon can only appear once in the dataset.
Slide 5 - Occurrence data
Other datasets published through GBIF.org have sufficiently consistent detail to contribute information about the location of individual organisms in time and space—that is, they offer evidence of the occurrence of a species (or other taxon) at a particular place on a specified date.
Occurrence datasets make up the core of data published through GBIF.org, and examples can range from specimens and fossils in natural history collections, observations by field researchers and citizen scientists, and data gathered from camera traps or remote-sensing satellites.
Occurrence records in these datasets sometimes provide only general locality information, sometimes simply identifying the country, but in many cases more precise locations and geographic coordinates support fine-scale analysis and mapping of species distributions.

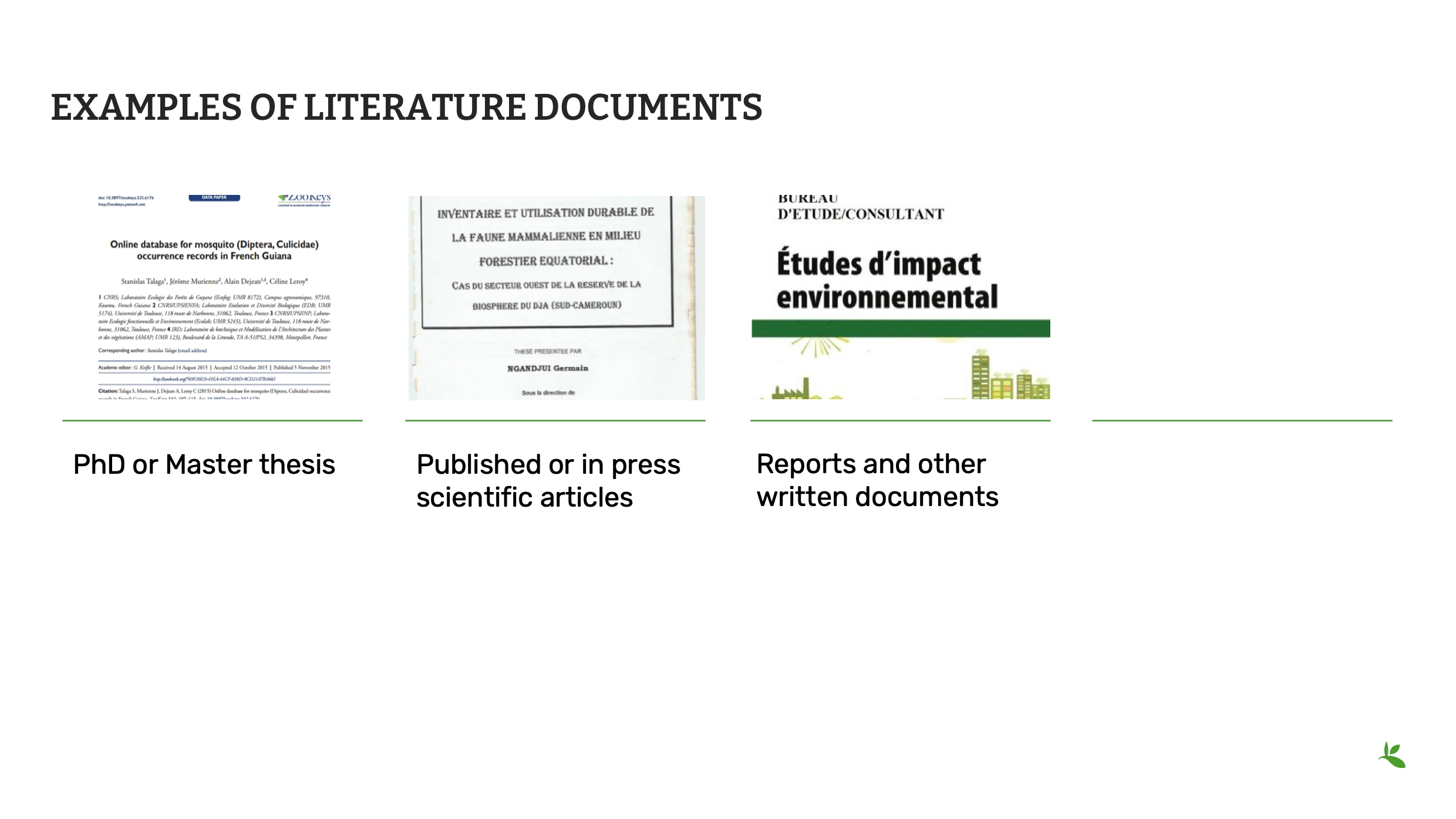
Slide 6 - Occurences from literature
Biodiversity data can also be found in technical and scientific literature : it is possible to compile and share this kind of data within GBIF, but you should be extra-cautious not to share duplicate data (for example, a specimen described in a scientific article might already be present on GBIF.org in the dataset of its collection)
En l’absence d’ensembles de données numérisés, les données sur la biodiversité peuvent être extraites et compilées à partir d’articles scientifiques, de thèses de doctorat ou de master, de rapports et d’autres documents. Contactez TOUJOURS le propriétaire de données d’abord lors de la compilation de données littéraires pour demander l’autorisation de les publier sur GBIF.

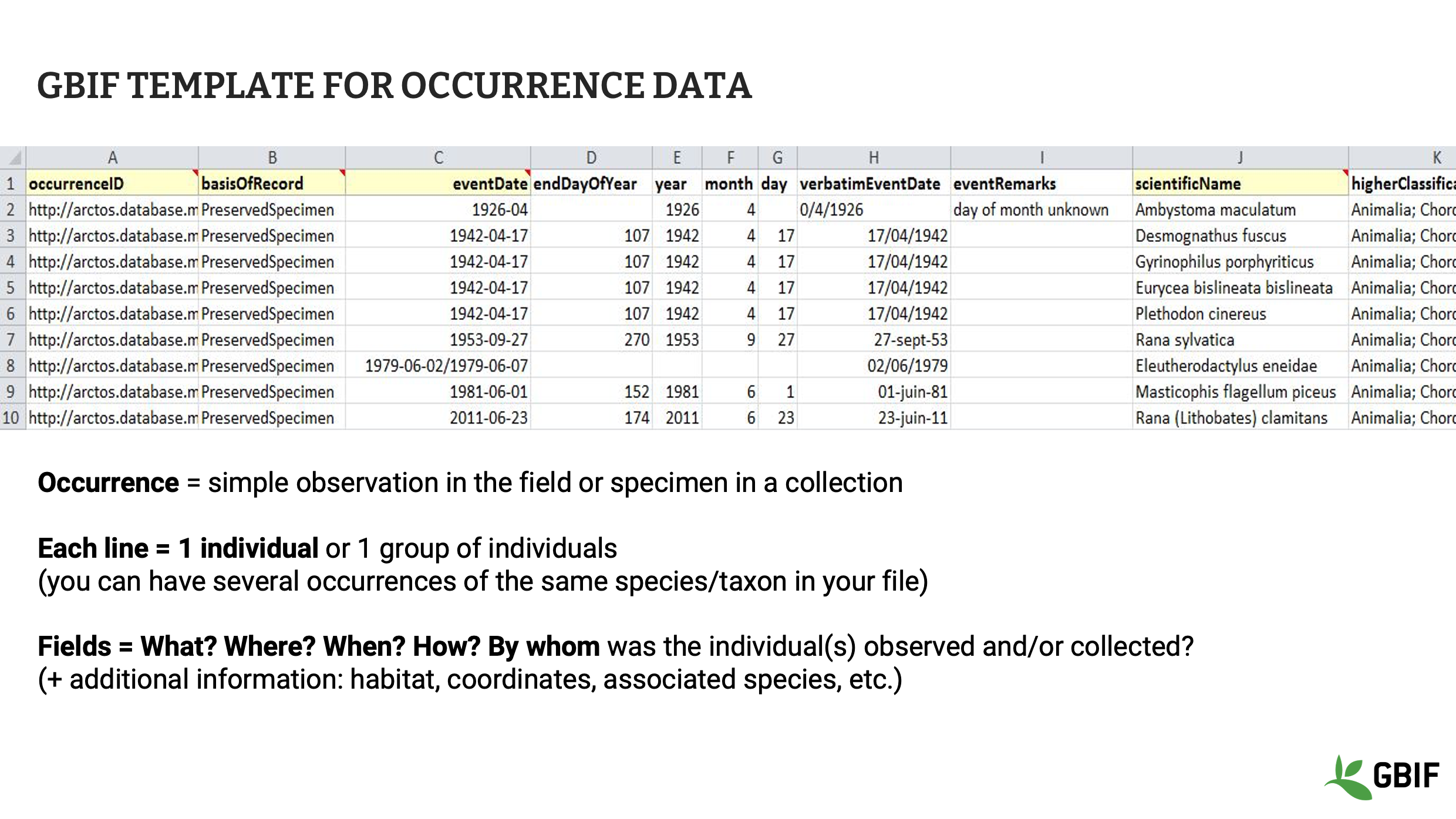
Slide 7 - GBIF template for occurrence data
Occurrence data represents the GBIF class of datasets with the largest number of records on GBIF.org. Collections, observations and literature data can be shared within GBIF using this template, which focuses on the observed individuals or collected specimens. In this kind of dataset, multiple individuals or specimens can be recorded for a single taxon, as long as they each have a unique identifier. Other fields for occurrence data include where, when, how and by whom was each occurrence observed and/or collected in the field.

Slide 8 - Sampling-event data
Datasets sometimes provide greater detail, not only offering evidence that a species occurred at a given location and date, but also making it possible to assess community composition for broader taxonomic groups or even the abundance of species at multiple times and places. These quantitative or sampling-event datasets typically derive from standard protocols for measuring and monitoring biodiversity like vegetation transects, bird censuses and freshwater or marine sampling.
By indicating the methods, events and relative abundance of species recorded in a sample, these datasets improve comparisons with data collected using the same protocols at different times and places—in some cases, even leading researchers to infer the absence of particular species from particular sites.

Slide 9 - GBIF template for sampling-event data
The GBIF template for Event data allows data publishers to share more information about the context of a biodiversity data collecting/recording event such as camera traps, insect traps, botanical relevés, birding sites and so on. Its structure is a more complex than the Taxonomical datasets and Occurrence datasets as it involves at least two files (e.g., two tabs in a spreadsheet) : one for describing the ‘events’ (e.g. each trap) and the other one to describe the specimens or occurrences linked to each event. There may be additional files/tabs to describe measurements or facts or to supply more details regarding sampling protocols.

Slide 10 - Dataset types and data quality requirements
GBIF provides data quality requirements to describe what you should provide for each dataset class. It doesn’t mean that the data won’t be indexed if some values are missing, but these requirements summarize what can be considered meaningful information for each class. You can of course share more.
Certaines exigences sont obligatoires (comme l’occurrenceID, le taxonID ou l’eventID, selon le type de jeu de données), d’autres sont facultatives mais fortement recommandées (comme les coordonnées en degrés décimaux). Vous trouverez ces exigences sur GBIF.org.

Slide 11 - How to choose a dataset type?
We know that choosing a dataset class can take some time, especially if you’re new to mobilizing data and sharing it with GBIF. The GBIF helpdesk team created this useful flowchart which can be found on the GBIF Data Blog if you need further support to decide.
Slide 12 - Reflection
In order to be able to share data with GBIF, it is important to understand the differences between the dataset classes at the early stages of the data capture and data managing process. You can reflect on this topic with the following questions:
Avec quel type de données travaillez-vous ?
Votre type de données est-il différent de ce que vous pensiez initialement ?
How would you publish them to GBIF?

Slide 13 - Conclusion
This is part of a series of presentations used in the GBIF Biodiversity Data Mobilization course. The biodiversity data mobilization curriculum was originally developed as part of the Biodiversity Information Development Programme funded by the European Union.
This presentation was originally created by Sophie Pamerlon with additional contributions by Laura Anne Russell and GBIF Trainers and Mentors. This presentation was narrated by me, Laura Anne Russell.
Saisie, traitement et qualité des données
|
In this video (12:06), you will explore the principles of data quality applied to data capture, specifically when capturing data from collection labels, fieldwork notebooks, spreadsheets, etc. If you are unable to watch the embedded video, you can download it locally. Subtitles are not yet available for this video. (MP4 - 42 MB) If you prefer to read, you will find the transcript below the embedded video. |
Transcription de la présentation
Cliquez pour développer

Diapositive 1 - Saisie, traitement et qualité des données
Cette présentation est basée sur les « Principes de qualité des données » de Arthur Chapman

Diapositive 2 - Structure
Au cours de cette présentation, nous allons explorer les principes de la qualité des données appliqués à la saisie des données, spécifiquement lors de la capture de données à partir des étiquettes de collecte, des cahiers de travail, des feuilles de calcul, etc.

Diapositive 3 - Développer un flux de travail pour le traitement et la qualité des données
La qualité des données est essentielle à chaque étape du processus de mobilisation des données, en particulier lors des étapes de saisie des données.
Chaque personne impliquée dans la saisie des données a sa part de responsabilité en ce qui concerne la qualité des données, mais la plupart des décisions à ce sujet doivent être prises au niveau institutionnel.
Les mots clés sont : planification et documentation !
Comme indiqué dans la présentation de la documentation sur les fondements, utilisez les normes existantes et planifiez vos flux de travail en fonction de vos objectifs ; documentez tout ce que vous pouvez, à chaque étape, et partagez ou réutilisez autant que possible les documents, les données, les outils et les normes.
Diapositive 4 - Développer un flux de travail pour le traitement et la qualité des données
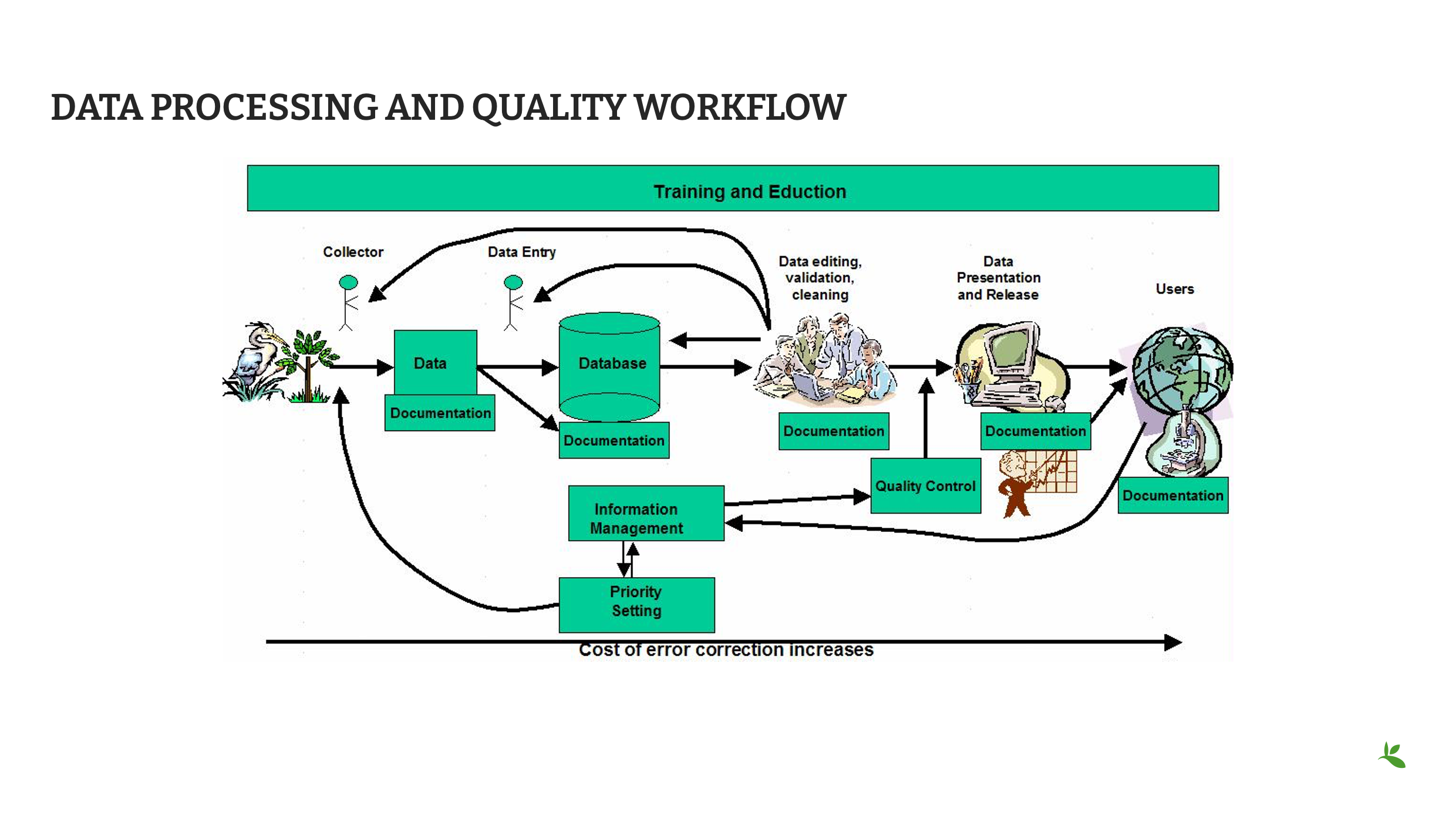
Il s’agit d’un exemple de flux de travail relatif à la qualité des données.
Ce flux de travail commence par la collecte des spécimens, passe à la saisie des données, puis au contrôle de la qualité, à la publication et enfin à l’utilisation.
La qualité des données n’est pas la seule responsabilité de la première personne du processus (ici, le collecteur) - elle est partagée à chaque étape et chaque personne du processus devrait être responsable de la qualité.
Une boucle de rétroaction fonctionnelle doit être mise en place pour vérifier, compléter, mettre à jour ou corriger les données.
C’est là que la documentation est essentielle : vous devez savoir qui était responsable de chaque étape du processus afin de valider les changements qui ont été apportés aux données (ou qui doivent être apportés aux données).

Diapositive 5 - Développer un flux de travail pour le traitement et la qualité des données
Dans cette vue simplifiée du flux de données, vous pouvez voir certaines des responsabilités en matière de qualité des données de chaque groupe de personnes impliquées.
Dans cet exemple, l’équipe chargée de la mobilisation peut être divisée entre les rôles de "transcripteurs" et de "conservateurs".
L’équipe de transcripteurs doit s’assurer que les données sont saisies et sauvegardées le mieux possible, tandis que le "conservateur" a la responsabilité ultime de veiller à ce que chaque équipe remplisse son rôle dans le processus.
Image du collectionneur : https://pixabay.com/fr/photos/d%C3%A9tective-loupe-regarde-un-788592/
Image de l’utilisateur : www.gbif.org (Résultat de la recherche du jeu de données WSC Cambodia camera trap dataset)
Photos du conservateur : https://pixabay.com/fr/photos/livres-livres-anciens-ordinateur-4802412/ & https://pixabay.com/fr/photos/donn%C3%A9es-ordinateur-internet-2899901/

Diapositive 6 - Structure
Une fois le flux de données mis en place, la saisie des données proprement dite peut commencer.
Dans les diapositives suivantes, nous allons explorer les différents types d’informations qui peuvent être obtenues à partir de spécimens ou d’observations sur le terrain, et nous verrons quelles sont les erreurs les plus courantes à éviter lorsque l’on traite chaque type d’information.
Les principaux thèmes abordés seront les suivants : informations taxonomiques, informations spatiales, informations sur les collections, informations descriptives.
Veuillez noter que chaque occurrence (chaque ligne de votre base de données ou de votre feuille de calcul) doit contenir des informations liées à ces quatre thèmes principaux afin d’être partagée et réutilisée en conséquence.

Diapositive 7 - Information taxonomique : vocabulaires et concepts
L’information taxonomique est un élément essentiel du processus de saisie des données.
Sans information taxonomique, un spécimen numérisé est inutile et ne peut être correctement interprété ou réutilisé.
Il convient de noter que le nom de l’espèce n’est pas le seul type d’information taxonomique pouvant être exploité dans le processus de saisie des données : parfois, le spécimen n’a pas été identifié jusqu’à l’espèce, et des niveaux taxonomiques plus élevés, tels que le genre ou la famille, sont toujours utiles pour les gestionnaires et les utilisateurs de données.

Diapositive 8 - Informations taxonomiques — attention aux noms!
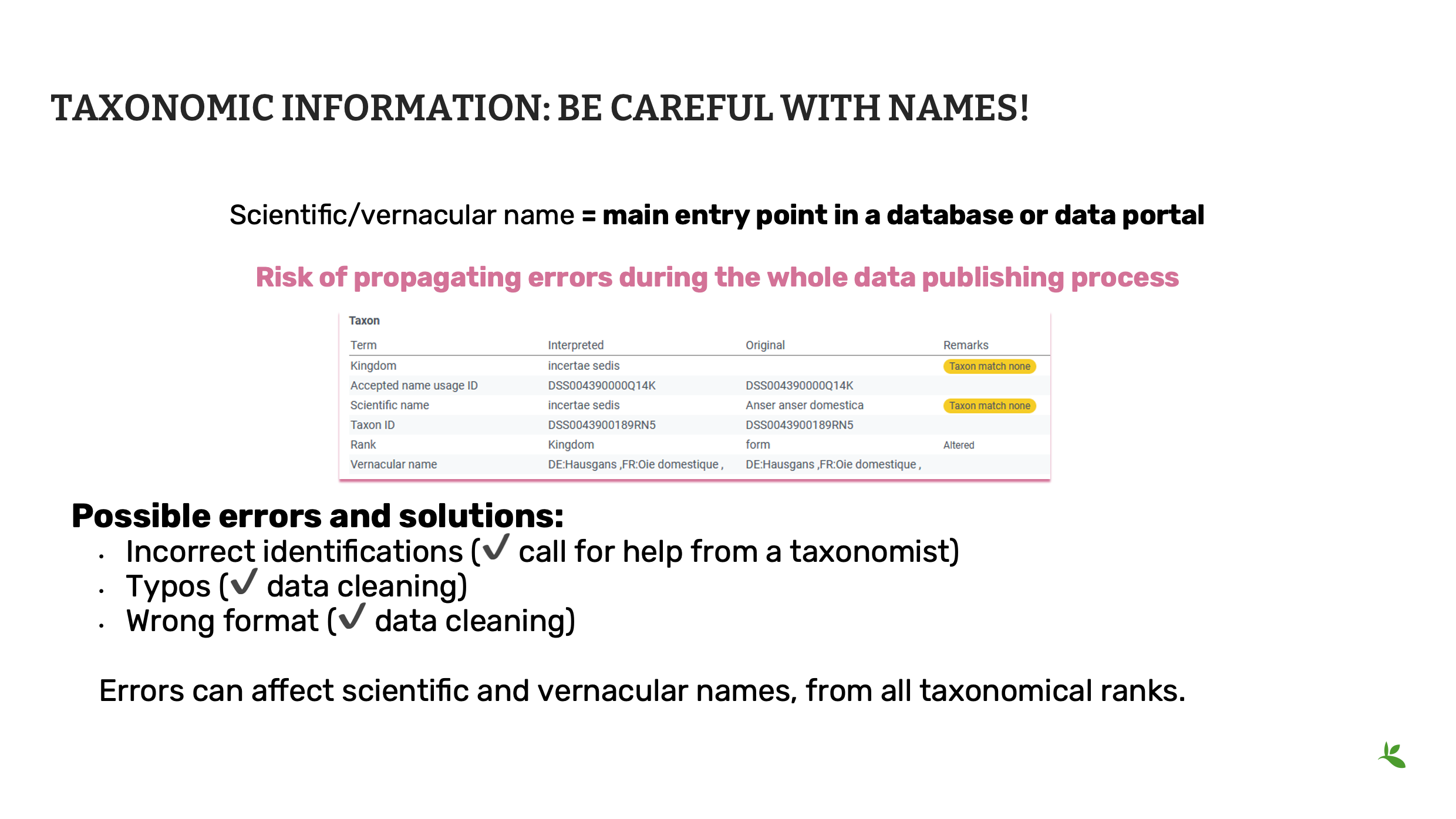
La plupart du temps, le nom scientifique est le principal moyen de retrouver des données dans une base de données, un portail, un site web, un navigateur, etc.
Toute erreur d’orthographe ou d’autorité peut conduire à des requêtes erronées ou nulles, entravant ainsi la gestion et la réutilisation potentielle des données.
C’est pourquoi il est très important de vérifier toutes les catégories de noms scientifiques afin de corriger les erreurs et/ou les omissions.

Diapositive 9 - Informations taxonomiques : erreurs communes à éviter
Les problèmes les plus courants concernant les informations sur les taxons sont les informations manquantes ou incohérentes, les valeurs incorrectes ou non atomiques, les doublons et l’incertitude.
Vérifiez toujours les définitions et les exemples des termes taxonomiques du Darwin Core pour éviter les erreurs de nomenclature : http://rs.tdwg.org/dwc/terms/index.htm

Diapositive 10 - Information taxonomique : vocabulaires et concepts
Les informations géographiques s’avèrent précieuses dans de nombreux contextes de réutilisation des données, tels que la modélisation des niches ou les études sur la répartition des espèces.
Bien qu’il soit difficile, voire impossible, de géolocaliser avec précision les collections ou les spécimens « anciens », il est recommandé de partager des coordonnées précises ou des informations textuelles lorsque cela est possible.
Les coordonnées doivent être enregistrées directement sur le terrain lorsque cela est possible, avec l’incertitude et le système de référence géodésique utilisé. Dans le cas contraire, utilisez des sources pertinentes et vérifiées pour géolocaliser vos données.
Il convient de noter que les coordonnées ou autres informations géographiques peuvent être généralisées ou ne pas être partagées du tout dans certains contextes, par exemple dans le cadre de la conservation d’espèces sensibles.

Diapositive 11 - Information spatiale : de quoi parlons-nous ?
Les informations spatiales peuvent se présenter sous de nombreux formats, et pas seulement sous la forme de coordonnées géographiques : les exemples incluent (mais ne sont pas limités à) des données de grille, des points+rayons ou des polygones.
Chacun d’entre eux est utile à partager afin de vérifier la cohérence des éléments géographiques (par exemple les coordonnées par rapport au code du pays, ou pour s’assurer qu’une localité donnée est cohérente avec les voyages d’un collectionneur).
Image de Point avec rayon : https://live.staticflickr.com/5820/23736048306_35dfd89c05_b.jpg

Diapositive 12 - Information spatiale : quelques définitions supplémentaires
Au sein du GBIF, il est recommandé de partager le système de référence géodésique utilisé pour dériver les coordonnées partagées (latitude et longitude décimales).
En l’absence d’un datum géodésique spécifique, GBIF déduira WGS84 par défaut.

Diapositive 13 - Informations taxonomiques : erreurs communes à éviter
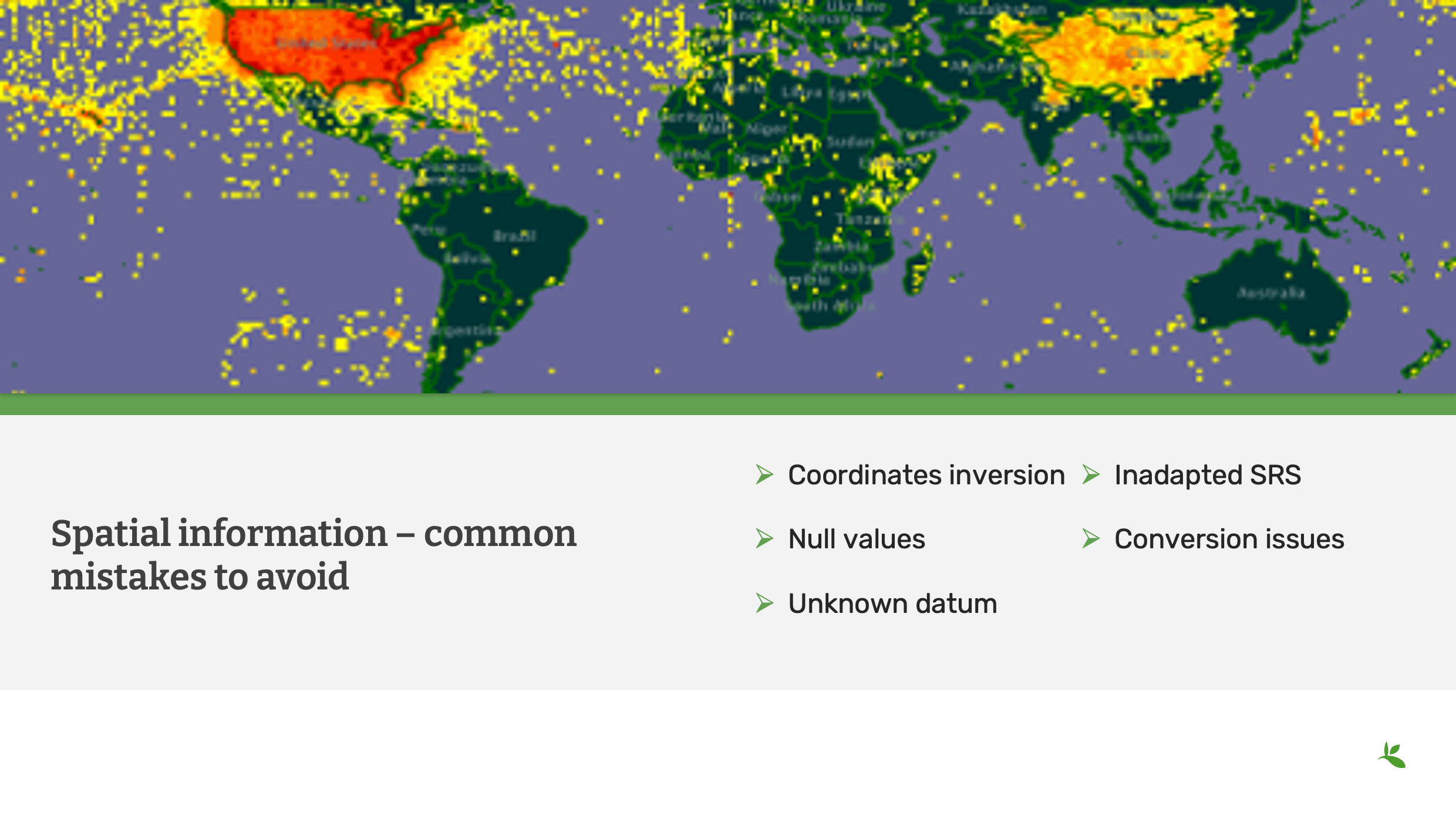
Cette diapositive montre une ancienne carte du GBIF présentant différents types de problèmes géographiques : le plus évident est un effet miroir entre les États-Unis et la Chine (coordonnées inversées),
Vous pouvez également remarquer une ligne artificielle le long du méridien de Greenwich où des valeurs '0' ont été mises dans le champ 'decimalLongitude', ainsi qu’une autre sur l’Équateur où des valeurs '0' ont été mises dans le champ 'decimalLatitude'.
L’indexation GBIF inclut désormais des vérifications géographiques automatiques entre les coordonnées et le code pays partagés dans le jeu de données. Les coordonnées peuvent être automatiquement inversées pour correspondre au pays.

Diapositive 14 - Information taxonomique : vocabulaires et concepts
Les informations sur le contexte de la collecte de données ou de l’observation sont très utiles à partager afin de donner autant de détails que possible sur chaque événement.
Des informations telles que le nom du collecteur, le protocole de collecte ou d’observation, l’habitat et d’autres facteurs peuvent s’avérer importantes lors de la réutilisation des données, par exemple dans le cadre de la modélisation des niches écologiques.
Selon le type d’ensemble de données, d’autres informations peuvent également s’avérer pertinentes.

Diapositive 15 - Informations sur la collecte : ce qu’il faut garder à l’esprit
Les facteurs de qualité des données concernant les informations collectées sont principalement l’exactitude, comme le nom correct du collecteur, la cohérence, par exemple l’utilisation du même vocabulaire pour décrire les sols et les habitats, et l’exhaustivité, comme la fourniture de toutes les informations existantes sur la description d’une espèce donnée, y compris la période de floraison, la couleur des feuilles et les utilisations médicinales.
Dans le Darwin Core et dans l’IPT, vous trouverez des vocabulaires contrôlés recommandés pour certains domaines tels que le "lifeStage". Le groupe de travail sur les vocabulaires du TDWG s’efforce de promouvoir et d’améliorer la facilité d’utilisation des vocabulaires.

Diapositive 16 - Information taxonomique : vocabulaires et concepts
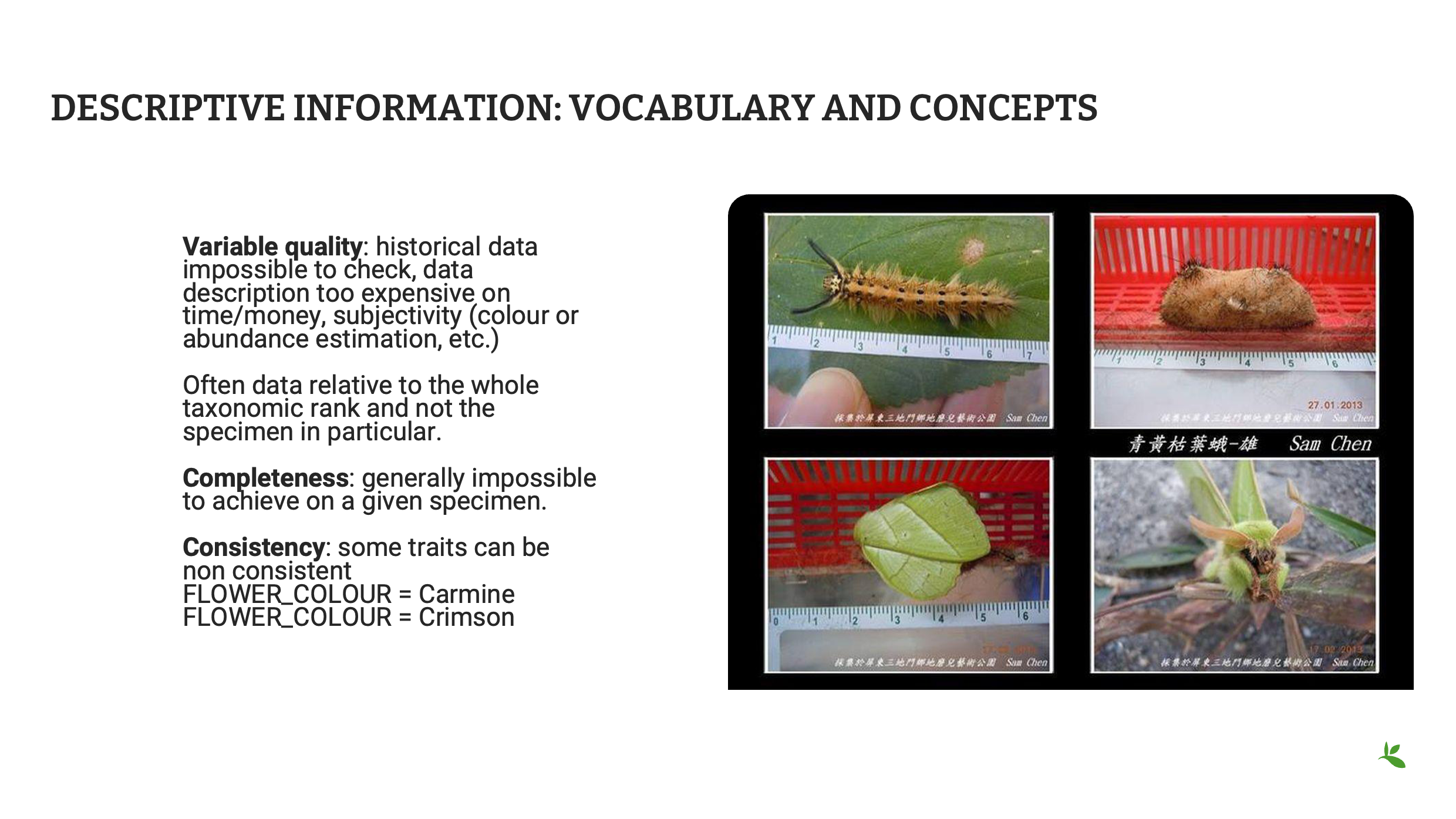
Gardez à l’esprit que les informations descriptives sont souvent incomplètes en raison d’une panoplie de facteurs.
Selon l’état de la collection, certaines étiquettes peuvent être incomplètes ou manquer d’informations essentielles ; l’exhaustivité (par exemple de la description d’une espèce) est souvent impossible à atteindre avec un seul individu ; et vous devez toujours vérifier la cohérence de votre base de données ou de votre feuille de calcul, par exemple dans les termes utilisés pour décrire les couleurs, afin d’éviter les informations redondantes.
Crédit image : Ensemble de données sur les occurrences de papillons de nuit à Taïwan recueillies sur les réseaux sociaux

Diapositive 17 - Résumé
Cette présentation s’est concentrée sur le thème de la qualité des données appliquée à la saisie des données ; en effet, il s’agit des étapes où il est crucial de s’assurer que toutes les informations relatives à chaque enregistrement sont correctement et complètement saisies, afin que les données soient aussi claires et compréhensibles que possible pour les futurs utilisateurs.
Cela n’est possible que si des décisions cohérentes sont prises au niveau institutionnel afin de créer un flux de travail solide pour la saisie et la gestion des données.
La chaîne de responsabilité concernant la qualité des données est alors répartie entre les personnes impliquées à chaque étape du processus, mais n’oubliez pas que les données peuvent toujours être améliorées et corrigées si des erreurs ou des omissions sont détectées à des stades ultérieurs.

Diapositive 18 - Conclusion
This is part of a series of presentations used in the GBIF Biodiversity Data Mobilization course. The biodiversity data mobilization curriculum was originally developed as part of the Biodiversity Information Development Programme funded by the European Union.
Cette présentation a été créée et narrée par Sophie Pamerlon avec des contributions supplémentaires des Coachs du BID et du BIFA, Mentors et Étudiants. La narration est par moi, Lily Shrestha.
Exercice 1c
|
Pour cette activité, vous réaliserez un exercice simulant la capture de données de l’analogique au numérique. Utilisez le site Darwin Core terms pour vous aider à prendre des décisions sur les données complémentaires nécessaires au projet et sur celles qui pourraient être partagées ultérieurement dans le cadre d’une publication. |
-
Télécharger UC-Practice-1-ForCapture-logs.zip. (939 KB). Le fichier compressé contient trois fichiers journaux de bord.
-
Téléchargez le modèle de feuille de calcul : UC-Practice-1-ForCapture-template.xlsx (27 KB) pour transcrire les observations enregistrées.
-
Utilisez la feuille d’exercice précédemment téléchargée pour donner vos réponses.
| vous devrez peut-être ajouter des champs à la feuille de calcul, car vous pourrez peut-être capturer plus d’informations à partir des étiquettes que ce qui était prévu dans le modèle. |